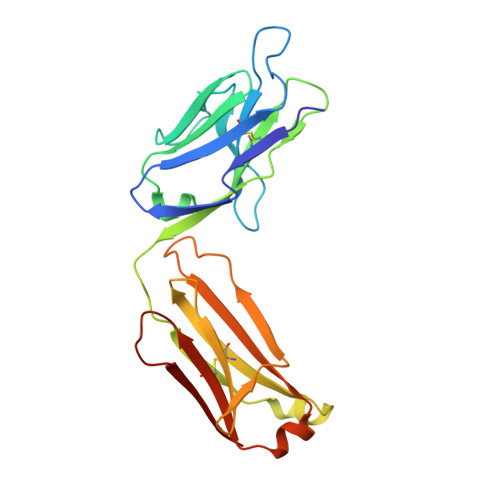
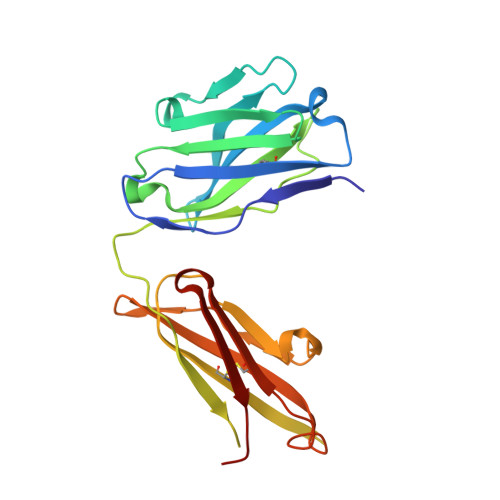
Crystal structure of an anticholera toxin peptide complex at 2.3 A.
Shoham, M.(1993) J Mol Biology 232: 1169-1175
- PubMed: 7690406 Search on PubMed
- DOI: https://doi.org/10.1006/jmbi.1993.1469
- Primary Citation Related Structures:
1TET - PubMed Abstract:
Cholera toxin peptide 3 (CTP3) is a 15-residue peptide corresponding in sequence to an immunogenic loop on the surface of the B-subunits of both cholera toxin and the heat-labile toxin from Escherichia coli. TE33 is the Fab fragment of a monoclonal antibody elicited against CTP3. The crystal structure of the TE33-CTP3 complex at 2.3 A resolution reveals an antigen-binding pocket, 13 A deep and 13 A wide, which is lined with many aromatic residues. The N-terminal portion of the peptide antigen CTP3 forms a type II beta-turn that fits snugly into this pocket. At gln7 the peptide backbone of CTP3 forms a kink followed by an extended C-terminal chain that seals off the cleft and buries the beta-turn underneath it. All six complementarity-determining regions of TE33 contribute to the binding of CTP3. The antibody-peptide contacts include, in addition to van der Waals' interactions and hydrogen bonds, also one salt bridge and one water molecule, which mediates the interaction.
- Case Western Reserve University, School of Medicine, Department of Biochemistry, Cleveland, OH 44106-4935.
Organizational Affiliation: