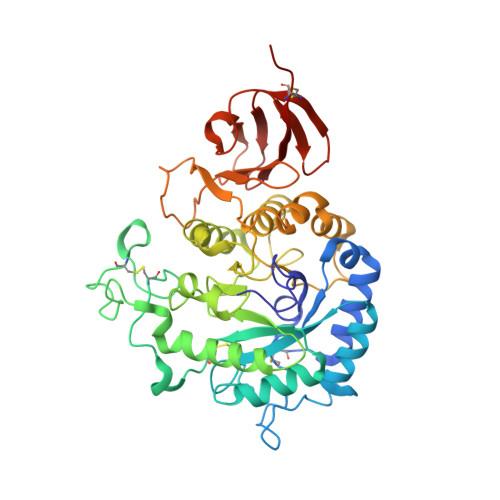
Crystal structure of alpha-galactosidase from Trichoderma reesei and its complex with galactose: implications for catalytic mechanism
Golubev, A.M., Nagem, R.A.P., Brandao Neto, J.R., Neustroev, K.N., Eneyskaya, E.V., Kulminskaya, A.A., Shabalin, K.A., Savel'ev, A.N., Polikarpov, I.(2004) J Mol Biology 339: 413-422
- PubMed: 15136043 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2004.03.062
- Primary Citation Related Structures:
1SZN, 1T0O - PubMed Abstract:
The crystal structures of alpha-galactosidase from the mesophilic fungus Trichoderma reesei and its complex with the competitive inhibitor, beta-d-galactose, have been determined at 1.54 A and 2.0 A resolution, respectively. The alpha-galactosidase structure was solved by the quick cryo-soaking method using a single Cs derivative. The refined crystallographic model of the alpha-galactosidase consists of two domains, an N-terminal catalytic domain of the (beta/alpha)8 barrel topology and a C-terminal domain which is formed by an antiparallel beta-structure. The protein contains four N-glycosylation sites located in the catalytic domain. Some of the oligosaccharides were found to participate in inter-domain contacts. The galactose molecule binds to the active site pocket located in the center of the barrel of the catalytic domain. Analysis of the alpha-galactosidase- galactose complex reveals the residues of the active site and offers a structural basis for identification of the putative mechanism of the enzymatic reaction. The structure of the alpha-galactosidase closely resembles those of the glycoside hydrolase family 27. The conservation of two catalytic Asp residues, identified for this family, is consistent with a double-displacement reaction mechanism for the alpha-galactosidase. Modeling of possible substrates into the active site reveals specific hydrogen bonds and hydrophobic interactions that could explain peculiarities of the enzyme kinetics.
- Petersburg Nuclear Physics Institute, Gatchina, St Petersburg, 188300, Russia.
Organizational Affiliation: