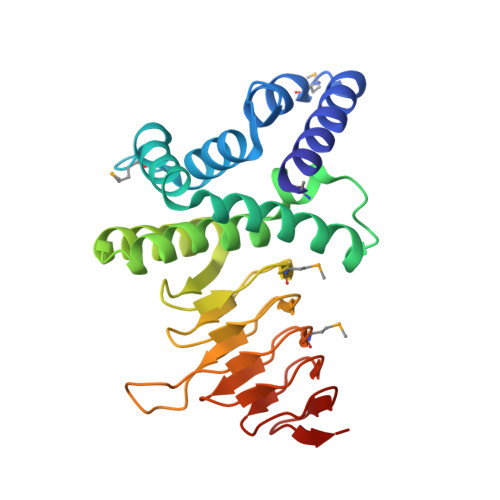
Structure of Serine Acetyltransferase in Complexes with CoA and its Cysteine Feedback Inhibitor
Olsen, L.R., Huang, B., Vetting, M.W., Roderick, S.L.(2004) Biochemistry 43: 6013-6019
- PubMed: 15147185 Search on PubMed
- DOI: https://doi.org/10.1021/bi0358521
- Primary Citation Related Structures:
1SSM, 1SSQ, 1SST - PubMed Abstract:
Serine acetyltransferase (SAT, EC 2.3.1.30) catalyzes the CoA-dependent acetylation of the side chain hydroxyl group of l-serine to form O-acetylserine, as the first step of a two-step biosynthetic pathway in bacteria and plants leading to the formation of l-cysteine. This reaction represents a key metabolic point of regulation for the cysteine biosynthetic pathway due to its feedback inhibition by cysteine. We have determined the X-ray crystal structure of Haemophilus influenzae SAT in complexes with CoA and its cysteine feedback inhibitor. The enzyme is a 175 kDa hexamer displaying the characteristic left-handed parallel beta-helix (LbetaH) structural domain of the hexapeptide acyltransferase superfamily of enzymes. Cysteine is bound in a crevice between adjacent LbetaH domains and underneath a loop excluded from the coiled LbetaH. The proximity of its thiol group to the thiol group of CoA derived from superimposed models of the cysteine and CoA complexes confirms that cysteine is bound at the active site. Analysis of the contacts of SAT with cysteine and CoA and the conformational differences that distinguish these complexes provides a structural basis for cysteine feedback inhibition, which invokes competition between cysteine and serine binding and a cysteine-induced conformational change of the C-terminal segment of the enzyme that excludes binding of the cofactor.
- Department of Biochemistry, Albert Einstein College of Medicine, 1300 Morris Park Avenue, Bronx, New York 10461, USA.
Organizational Affiliation: