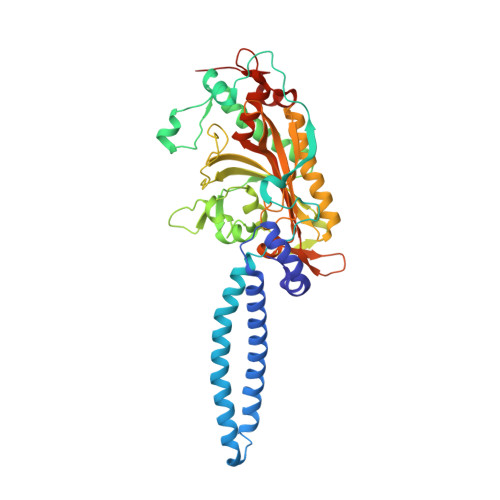
Refined crystal structure of the seryl-tRNA synthetase from Thermus thermophilus at 2.5 A resolution.
Fujinaga, M., Berthet-Colominas, C., Yaremchuk, A.D., Tukalo, M.A., Cusack, S.(1993) J Mol Biology 234: 222-233
- PubMed: 8230201 Search on PubMed
- DOI: https://doi.org/10.1006/jmbi.1993.1576
- Primary Citation Related Structures:
1SRY - PubMed Abstract:
The three-dimensional structure of the seryl-tRNA synthetase from Thermus thermophilus has been determined and refined at 2.5 A resolution. The final model consists of a dimer of 421 residues each and 190 water molecules. The R-factor is 18.4% for all the data between 10 and 2.5 A resolution. The structure is very similar to that of the homologous enzyme from Escherichia coli, with an r.m.s. difference of 1.5 A for the 357 alpha-carbon atoms considered equivalent. The comparison of the two structures indicates increased hydrophobicity, reduced conformational entropy and reduced torsional strain as possible mechanisms by which thermostability is obtained in the enzyme from the thermophile.
- EMBL Grenoble Outstation, France.
Organizational Affiliation: