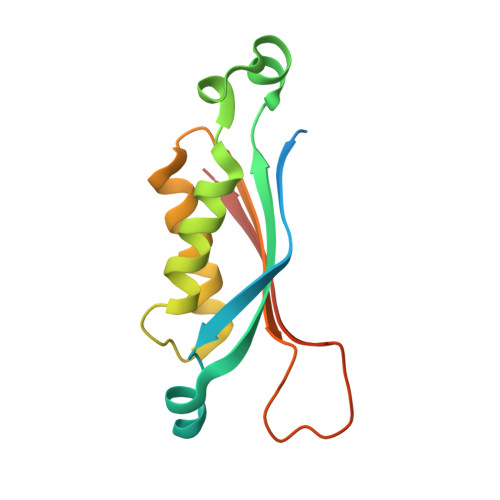
Biosynthesis of Tetrahydrofolate in Plants: Crystal Structure of 7,8-Dihydroneopterin Aldolase from Arabidopsis thaliana Reveals a Novel Adolase Class.
Bauer, S., Schott, A.K., Illarionova, V., Bacher, A., Huber, R., Fischer, M.(2004) J Mol Biology 339: 967-979
- PubMed: 15165863 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2004.04.034
- Primary Citation Related Structures:
1SQL - PubMed Abstract:
Dihydroneopterin aldolase (DHNA) catalyses a retroaldol reaction yielding 6-hydroxymethyl-7,8-dihydropterin, a biosynthetic precursor of the vitamin, tetrahydrofolate. The enzyme is a potential target for antimicrobial and anti-parasite chemotherapy. A gene specifying a dihydroneopterin aldolase from Arabidopsis thaliana was expressed in a recombinant Escherichia coli strain. The recombinant protein was purified to apparent homogeneity and crystallised using polyethylenglycol as the precipitating agent. The crystal structure was solved by X-ray diffraction analysis at 2.2A resolution. The enzyme forms a D(4)-symmetric homooctamer. Each polypeptide chain is folded into a single domain comprising an antiparallel four-stranded beta-sheet and two long alpha-helices. Four monomers are arranged in a tetrameric ring, and two of these rings form a hollow cylinder. Well defined purine derivatives are found at all eight topologically equivalent active sites. The subunit fold of the enzyme is related to substructures of dihydroneopterin triphosphate epimerase, GTP cyclohydrolase I, and pyruvoyltetrahydropterin synthase, which are all involved in the biosynthesis of pteridine type cofactors, and to urate oxidase, although some members of that superfamily have no detectable sequence similarity. Due to structural and mechanistical differences of DHNA in comparison with class I and class II aldolases, a new aldolase class is proposed.
- Max-Planck-Institut für Biochemie, Abteilung Strukturforschung, Am Klopferspitz 18a, D-82152 Martinsried, Germany. stbauer@biochem.mpg.de
Organizational Affiliation: