Product of enzymatic self-cleavage bound in the active site of HIV protease
Vondrackova, E., Hasek, J., Jaskolski, M., Rezacova, P., Dohnalek, J., Skalova, T., Petrokova, H., Duskova, J., Brynda, J., Sedlacek, J.To be published.
Experimental Data Snapshot
Starting Model: experimental
View more details
wwPDB Validation 3D Report Full Report
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Protease | 99 | Human immunodeficiency virus 1 | Mutation(s): 0 Gene Names: POL EC: 3.4.23.16 | 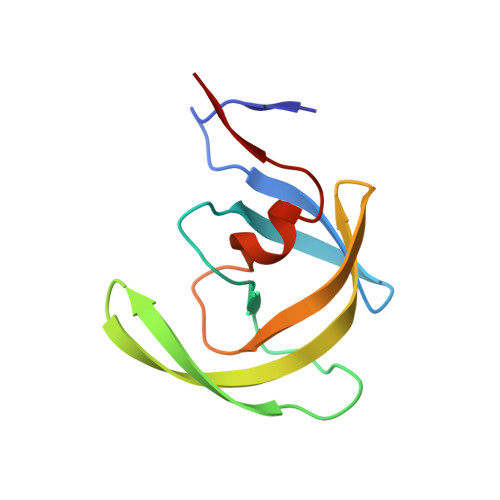 | |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P03367 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| 5-mer peptide from Protease | C [auth I] | 5 | Human immunodeficiency virus 1 | Mutation(s): 0 Gene Names: POL | 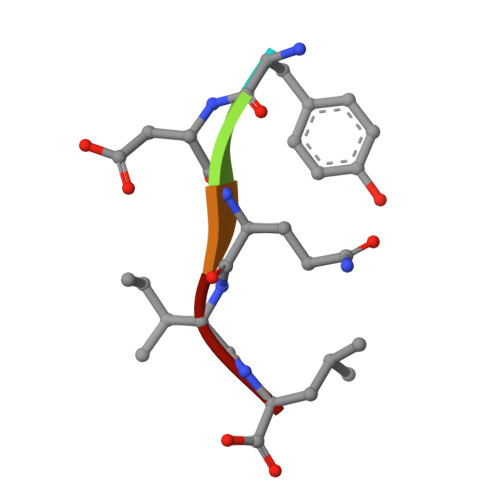 |
UniProt | |||||
Entity Groups | |||||
| UniProt Group | P03367 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
| Ligands 3 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| BME Download:Ideal Coordinates CCD File | F [auth A], G [auth A], J [auth B] | BETA-MERCAPTOETHANOL C2 H6 O S DGVVWUTYPXICAM-UHFFFAOYSA-N |  | ||
| DMS Download:Ideal Coordinates CCD File | H [auth A] | DIMETHYL SULFOXIDE C2 H6 O S IAZDPXIOMUYVGZ-UHFFFAOYSA-N |  | ||
| CL Download:Ideal Coordinates CCD File | D [auth A], E [auth A], I [auth B] | CHLORIDE ION Cl VEXZGXHMUGYJMC-UHFFFAOYSA-M |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 57.765 | α = 90 |
| b = 86.284 | β = 90 |
| c = 45.834 | γ = 90 |
| Software Name | Purpose |
|---|---|
| CNS | refinement |
| DENZO | data reduction |
| SCALEPACK | data scaling |
| CNS | phasing |