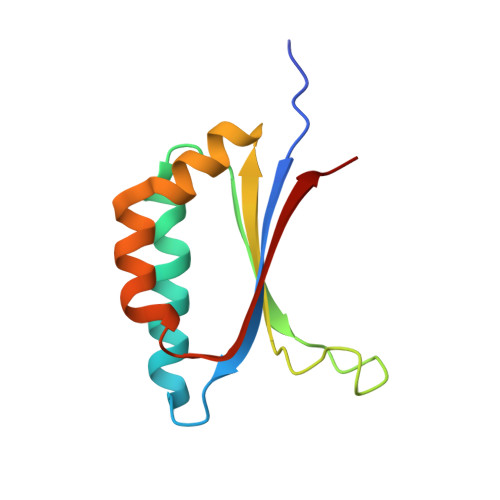
The Structural Basis of the Thermostability of SP1, a Novel Plant (Populus tremula) Boiling Stable Protein.
Dgany, O., Gonzalez, A., Sofer, O., Wang, W., Zolotnitsky, G., Wolf, A., Shoham, Y., Altman, A., Wolf, S.G., Shoseyov, O., Almog, O.(2004) J Biological Chem 279: 51516-51523
- PubMed: 15371455 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M409952200
- Primary Citation Related Structures:
1SI9, 1TR0 - PubMed Abstract:
We previously reported on a new boiling stable protein isolated from aspen plants (Populus tremula), which we named SP1. SP1 is a stress-related protein with no significant sequence homology to other stress-related proteins. It is a 108-amino-acid hydrophilic polypeptide with a molecular mass of 12.4 kDa (Wang, W. X., Pelah, D., Alergand, T., Shoseyov, O., and Altman, A. (2002) Plant Physiol. 130, 865-875) and is found in an oligomeric form. Preliminary electron microscopy studies and matrix-assisted laser desorption ionization time-of-flight mass spectrometry experiments showed that SP1 is a dodecamer composed of two stacking hexamers. We performed a SDS-PAGE analysis, a differential scanning calorimetric study, and crystal structure determination to further characterize SP1. SDS-PAGE indicated a spontaneous assembly of SP1 to one stable oligomeric form, a dodecamer. Differential scanning calorimetric showed that SP1 has high thermostability i.e. Tm of 107 degrees C (at pH 7.8). The crystal structure of SP1 was initially determined to 2.4 A resolution by multi-wavelength anomalous dispersion method from a crystal belonging to the space group I422. The phases were extended to 1.8 A resolution using data from a different crystal form (P21). The final refined molecule includes 106 of the 108 residues and 132 water molecules (on average for each chain). The R-free is 20.1%. The crystal structure indicated that the SP1 molecule has a ferredoxin-like fold. Strong interactions between each two molecules create a stable dimer. Six dimers associate to form a ring-like-shaped dodecamer strongly resembling the particle visualized in the electron microscopy studies. No structural similarity was found between the crystal structure of SP1 and the crystal structure of other stress-related proteins such as small heat shock proteins, whose structure has been already determined. This structural study further supports our previous report that SP1 may represent a new family of stress-related proteins with high thermostability and oligomerization.
- Department of Clinical Biochemistry, Faculty of Health Sciences, Ben-Gurion University, Beer-Sheva 84105, Israel.
Organizational Affiliation: