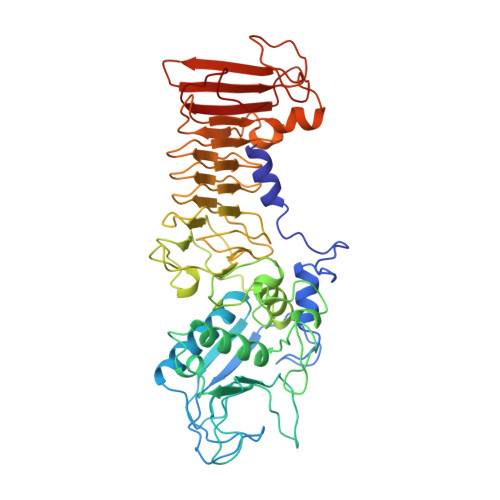
Crystal structure of the 50 kDa metallo protease from Serratia marcescens.
Baumann, U.(1994) J Mol Biology 242: 244-251
- PubMed: 8089845 Search on PubMed
- DOI: https://doi.org/10.1006/jmbi.1994.1576
- Primary Citation Related Structures:
1SAT - PubMed Abstract:
The crystal structure of the 50 kDa metalloprotease from the Gram-negative bacterium Serratia marcescens has been solved and refined to a crystallographic R-factor of 0.192 at 1.80 A resolution. The structure is very similar to that of alkaline protease from Pseudomonas aeruginosa, in particular the calcium binding "parallel beta roll" motif is completely conserved. The N-terminal proteolytic domain shows the typical "metzincin" fold. The active sites of the two enzymes are slightly different, Tyr216 is a Zn ligand in the Serratia metallo protease. The loops 70-77 and 122-132, which encompass the active site cleft, differ due to insertions and deletions so that the Serratia metallo protease seems to have a more open site than the alkaline protease.
- Abteilung für Strukturforschung, Max-Planck-Institut für Biochemie, Martinsried, Germany.
Organizational Affiliation: