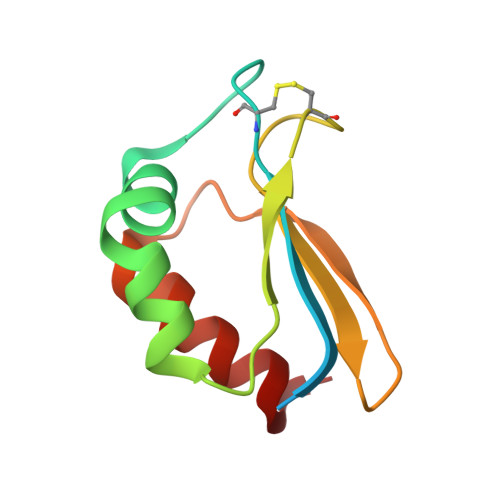
The oxidized subunit B8 from human complex I adopts a thioredoxin fold.
Brockmann, C., Diehl, A., Rehbein, K., Strauss, H., Schmieder, P., Korn, B., Kuhne, R., Oschkinat, H.(2004) Structure 12: 1645-1654
- PubMed: 15341729 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2004.06.021
- Primary Citation Related Structures:
1S3A - PubMed Abstract:
Subunit B8 from ubiquinone oxidoreductase (complex I) (CI-B8) is one of several nuclear-encoded supernumerary subunits that are not present in bacterial complex I. Its solution structure shows a thioredoxin fold with highest similarities to the human thioredoxin mutant C73S and thioredoxin 2 from Anabeana sp. Interestingly, these proteins contain active sites in the same area, where the disulfide bond of oxidized CI-B8 is located. The redox potential of this disulfide bond is -251.6 mV, comparing well to that of disulfides in other thioredoxin-like proteins. Analysis of the structure reveals a surface area that is exclusively composed of highly conserved residues and thus most likely a subunit interaction site within complex I.
- Forschungsinstitut für Molekulare Pharmakologie, D-13125 Berlin, Germany.
Organizational Affiliation: