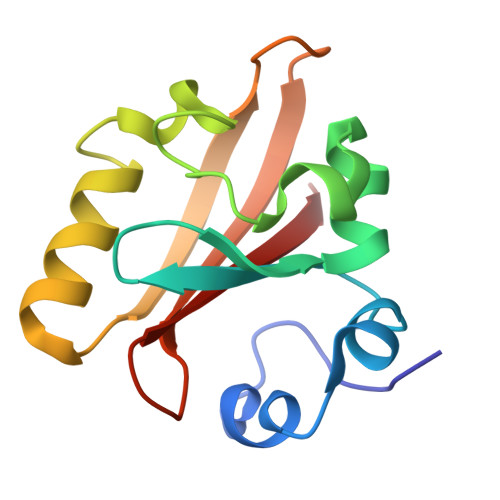
Chromophore conformation and the evolution of tertiary structural changes in photoactive yellow protein
Anderson, S., Srajer, V., Pahl, R., Rajagopal, S., Schotte, F., Anfinrud, P., Wulff, M., Moffat, K.(2004) Structure 12: 1039-1045
- PubMed: 15274923 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2004.04.008
- Primary Citation Related Structures:
1S1Y, 1S1Z - PubMed Abstract:
We use time-resolved crystallography to observe the structural progression of a bacterial blue light photoreceptor throughout its photocycle. Data were collected from 10 ns to 100 ms after photoactivation of the E46Q mutant of photoactive yellow protein. Refinement of transient chromophore conformations shows that the spectroscopically distinct intermediates are formed via progressive disruption of the hydrogen bond network to the chromophore. Although structural change occurs within a few nanoseconds on and around the chromophore, it takes milliseconds for a distinct pattern of tertiary structural change to fully progress through the entire molecule, thus generating the putative signaling state. Remarkably, the coupling between the chromophore conformation and the tertiary structure of this small protein is not tight: there are leads and lags between changes in the conformation of the chromophore and the protein tertiary structure.
- Department of Biochemistry and Molecular Biology, University of Chicago, Chicago, IL 60637, USA. smander@midway.uchicago.edu
Organizational Affiliation: