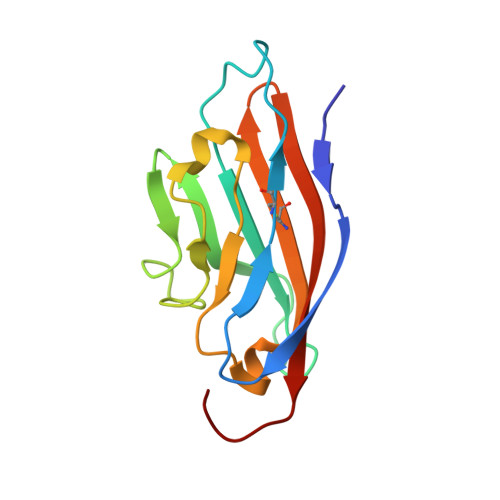
Solution structure of the coxsackievirus and adenovirus receptor domain 1.
Jiang, S., Jacobs, A., Laue, T.M., Caffrey, M.(2004) Biochemistry 43: 1847-1853
- PubMed: 14967025 Search on PubMed
- DOI: https://doi.org/10.1021/bi035490x
- Primary Citation Related Structures:
1RSF - PubMed Abstract:
The coxsackievirus and adenovirus receptor (CAR) mediates entry of coxsackievirus B (CVB) and adenovirus (Ad). The normal cellular function of CAR, which is expressed in a wide variety of tissue types, is thought to involve homophilic cell adhesion in the developing brain. The extracellular domain of CAR consists of two immunoglobulin (Ig) domains termed CAR-D1 and CAR-D2. CAR-D1 is shown by sedimentation velocity to be monomeric at pH 3.0. The solution structure and the dynamic properties of monomeric CAR-D1 have been determined by NMR spectroscopy at pH 3.0. The determinants of the CAR-D1 monomer-dimer equilibrium, as well as the binding site of CVB and Ad on CAR, are discussed in light of the monomer structure.
- Department of Biochemistry and Moleular Genetics, University of Illinois at Chicago, Chicago, Illinois 60607, USA.
Organizational Affiliation: