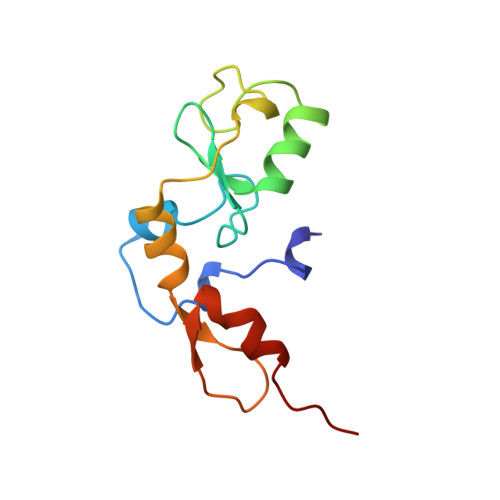
Crystal structure of the RAG1 dimerization domain reveals multiple zinc-binding motifs including a novel zinc binuclear cluster.
Bellon, S.F., Rodgers, K.K., Schatz, D.G., Coleman, J.E., Steitz, T.A.(1997) Nat Struct Biol 4: 586-591
- PubMed: 9228952 Search on PubMed
- DOI: https://doi.org/10.1038/nsb0797-586
- Primary Citation Related Structures:
1RMD - PubMed Abstract:
The crystal structure of the dimerization domain of the V(D)J recombination-activating protein, RAG1, was solved using zinc anomalous scattering. The structure reveals an unusual combination of multi-class zinc-binding motifs, including a zinc RING finger and a C2H2 zinc finger, that together from a single structural domain. The domain also contains a unique zinc binuclear cluster in place of a normally mononuclear zinc site in the RING finger. Together, four zinc ions help organize the entire domain, including the two helices that form the dimer interface.
- Department of Molecular Biophysics and Biochemistry, Yale University, New Haven, Connecticut 06520-8114, USA.
Organizational Affiliation: