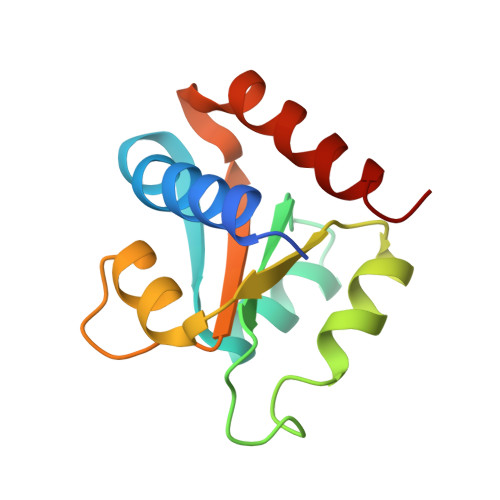
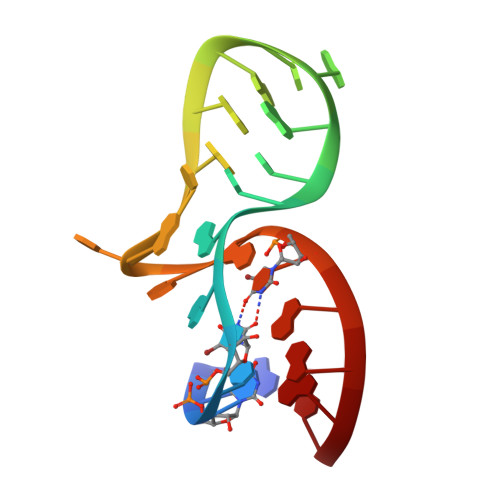
Molecular Basis of Box C/D RNA-Protein Interactions; Cocrystal Structure of Archaeal L7Ae and a Box C/D RNA.
Moore, T., Zhang, Y., Fenley, M.O., Li, H.(2004) Structure 12: 807-818
- PubMed: 15130473 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2004.02.033
- Primary Citation Related Structures:
1RLG - PubMed Abstract:
We have determined and refined a crystal structure of the initial assembly complex of archaeal box C/D sRNPs comprising the Archaeoglobus fulgidus (AF) L7Ae protein and a box C/D RNA. The box C/D RNA forms a classical kink-turn (K-turn) structure and the resulting protein-RNA complex serves as a distinct platform for recruitment of the fibrillarin-Nop5p complex. The cocrystal structure confirms previously proposed secondary structure of the box C/D RNA that includes a protruded U, a UU mismatch, and two sheared tandem GA base pairs. Detailed structural comparisons of the AF L7Ae-box C/D RNA complex with previously determined crystal structures of L7Ae homologs in complex with functionally distinct K-turn RNAs revealed a set of remarkably conserved principles in protein-RNA interactions. These analyses provide a structural basis for interpreting the functional roles of the box C/D sequences in directing specific assembly of box C/D sRNPs.
- Department of Chemistry and Biochemistry, Florida State University, Tallahassee, FL 32306, USA.
Organizational Affiliation: