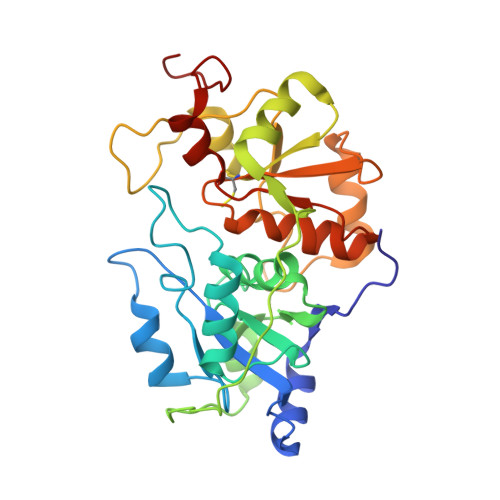
Structure of bovine liver rhodanese. I. Structure determination at 2.5 A resolution and a comparison of the conformation and sequence of its two domains.
Ploegman, J.H., Drent, G., Kalk, K.H., Hol, W.G.(1978) J Mol Biology 123: 557-594
- PubMed: 691057 Search on PubMed
- Primary Citation Related Structures:
1RHD