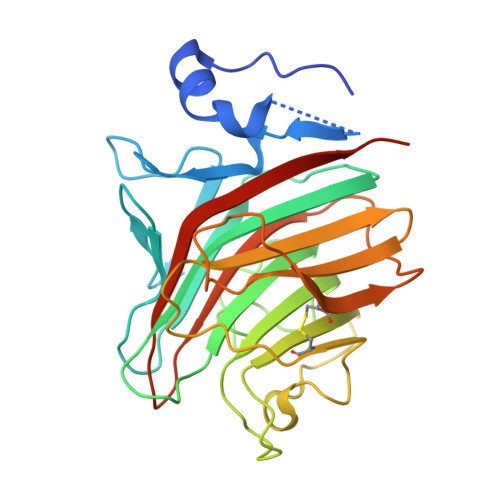
The Crystal Structure of the Carbohydrate-recognition Domain of the Glycoprotein Sorting Receptor p58/ERGIC-53 Reveals an Unpredicted Metal-binding Site and Conformational Changes Associated with Calcium Ion Binding.
Velloso, L.M., Svensson, K., Pettersson, R.F., Lindqvist, Y.(2003) J Mol Biology 334: 845-851
- PubMed: 14643651 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2003.10.031
- Primary Citation Related Structures:
1R1Z - PubMed Abstract:
p58/ERGIC-53 is a calcium-dependent animal lectin that acts as a cargo receptor, binding to a set of glycoproteins in the endoplasmic reticulum (ER) and transporting them to the Golgi complex. It is similar in structure to calcium-dependent leguminous lectins. We have determined the structure of the carbohydrate-recognition domain of p58/ERGIC-53 in its calcium-bound form. The structure reveals localized but large conformational changes in relation to the previously determined metal ion-free structure, mapping mostly to the ligand-binding site. It reveals the presence of two calcium ion-binding sites located 6A apart, one of which has no equivalent in the plant lectins. The second metal ion-binding site present in that class of lectins, binding Mn(2+), is absent from p58/ERGIC-53. The absence of a short loop in the ligand-binding site in this protein suggests that it has adapted to optimally bind the high-mannose Man(8)(GlcNAc)(2) glycan common to glycoproteins at the ER exit stage.
- Molecular Structural Biology, Department of Medical Biochemistry and Biophysics, Karolinska Institutet, S-171 77 Stockholm, Sweden.
Organizational Affiliation: