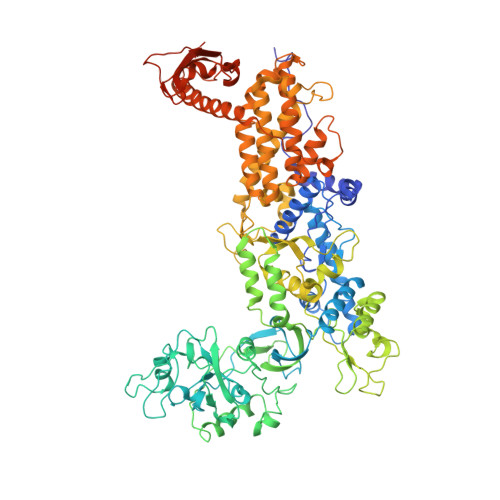
Insights into editing from an ile-tRNA synthetase structure with tRNAile and mupirocin.
Silvian, L.F., Wang, J., Steitz, T.A.(1999) Science 285: 1074-1077
- PubMed: 10446055 Search on PubMed
- Primary Citation Related Structures:
1FFY, 1QU2, 1QU3 - PubMed Abstract:
Isoleucyl-transfer RNA (tRNA) synthetase (IleRS) joins Ile to tRNA(Ile) at its synthetic active site and hydrolyzes incorrectly acylated amino acids at its editing active site. The 2.2 angstrom resolution crystal structure of Staphylococcus aureus IleRS complexed with tRNA(Ile) and Mupirocin shows the acceptor strand of the tRNA(Ile) in the continuously stacked, A-form conformation with the 3' terminal nucleotide in the editing active site. To position the 3' terminus in the synthetic active site, the acceptor strand must adopt the hairpinned conformation seen in tRNA(Gln) complexed with its synthetase. The amino acid editing activity of the IleRS may result from the incorrect products shuttling between the synthetic and editing active sites, which is reminiscent of the editing mechanism of DNA polymerases.
- Department of Molecular Biophysics, Yale University, and Howard Hughes Medical Institute, New Haven, CT 06520-8114, USA.
Organizational Affiliation: