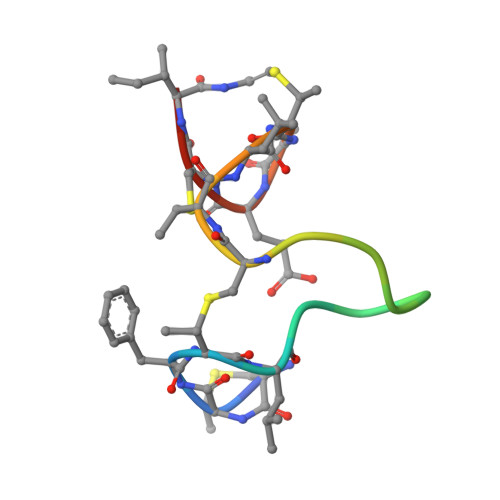
Ab initio structure determination of the lantibiotic mersacidin.
Schneider, T.R., Karcher, J., Pohl, E., Lubini, P., Sheldrick, G.M.(2000) Acta Crystallogr D Biol Crystallogr 56: 705-713
- PubMed: 10818347 Search on PubMed
- DOI: https://doi.org/10.1107/s0907444900003711
- Primary Citation Related Structures:
1QOW - PubMed Abstract:
The crystal structure of mersacidin, a potential novel antibiotic against methicillin- and vancomycin-resistant Staphylococcus aureus strains, has been determined by ab initio methods. Despite all crystals being merohedrally twinned, an accurate structural model with an R value of 13.4% has been obtained at atomic resolution. With six molecules in the asymmetric unit and no atom heavier than sulfur, the structure corresponds to a protein of 120 amino acids and is the largest approximately equal-atom unknown structure solved by direct methods. In the crystal, the molecule assumes a compact fold different from that found by NMR in solution. Comparison of the NCS-related molecules reveals regions of variable flexibility. The region highly homologous to the related antibiotic actagardine is very rigid and possibly defines an essential building block of this class of new antibacterial substances.
- University of Göttingen, Department of Structural Chemistry, Germany. trs@shelx.uni-ac-gwdg.de
Organizational Affiliation: