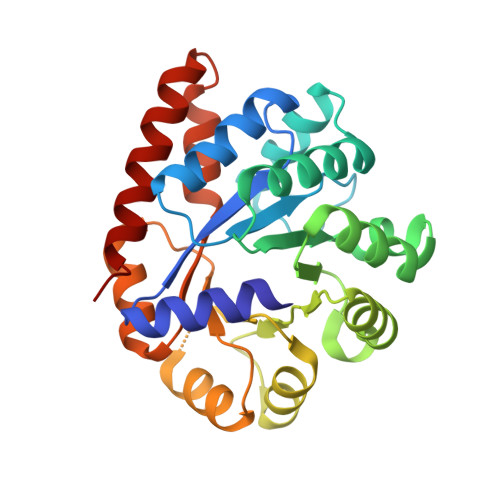
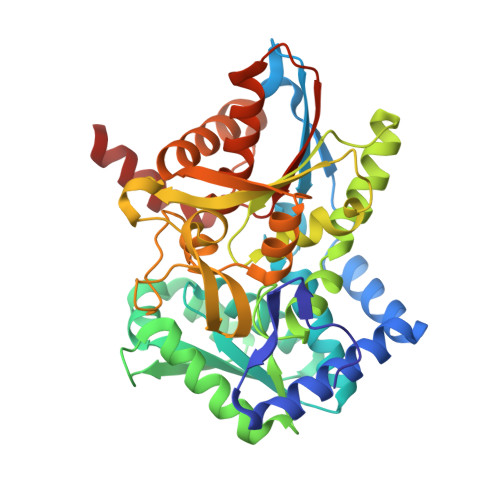
Crystal Structure of Wild-Type Tryptophan Synthase Complexed with the Natural Substrate Indole-3-Glycerol Phosphate.
Weyand, M., Schlichting, I.(1999) Biochemistry 38: 16469
- PubMed: 10600108 Search on PubMed
- DOI: https://doi.org/10.1021/bi9920533
- Primary Citation Related Structures:
1QOP, 1QOQ - PubMed Abstract:
We used freeze trapping to stabilize the Michaelis complex of wild-type tryptophan synthase and the alpha-subunit substrate indole-3-glycerol phosphate (IGP) and determined its structure to 1. 8 A resolution. In addition, we determined the 1.4 A resolution structure of the complex with indole-3-propanole phosphate (IPP), a noncleavable IGP analogue. The interaction of the 3'-hydroxyl of IGP with the catalytic alphaGlu49 leads to a twisting of the propane chain and to a repositioning of the indole ring compared to IPP. Concomitantly, the catalytic alphaAsp60 rotates resulting in a translocation of the COMM domain [betaGly102-betaGly189, for definition see Schneider et al. (1998) Biochemistry 37, 5394-5406] in a direction opposite to the one in the IPP complex. This results in loss of the allosteric sodium ion bound at the beta-subunit and an opening of the beta-active site, thereby making the cofactor pyridoxal 5'-phosphate (PLP) accessible to solvent and thus serine binding. These findings form the structural basis for the information transfer from the alpha- to the beta-subunit and may explain the affinity increase of the beta-active site for serine upon IGP binding.
- Max-Planck-Institute for Molecular Physiology, Department of Physical Biochemistry, Dortmund, Germany.
Organizational Affiliation: