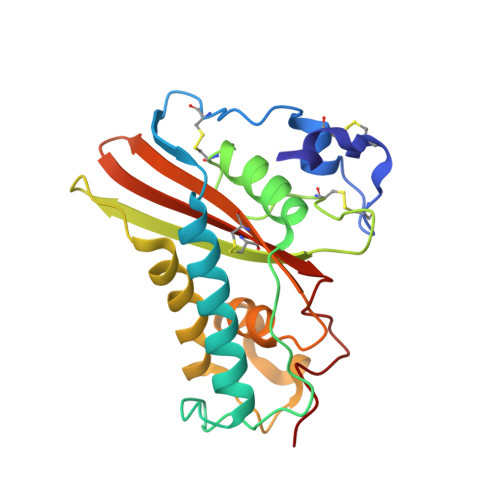
Major Venom Allergen of Yellow Jackets, Ves V 5: Structural Characterization of a Pathogenesis-Related Protein Superfamily.
Henriksen, A., King, T.P., Mirza, O., Monsalve, R.I., Meno, K., Ipsen, H., Larsen, J.N., Gajhede, M., Spangfort, M.D.(2001) Proteins 45: 438
- PubMed: 11746691 Search on PubMed
- DOI: https://doi.org/10.1002/prot.1160
- Primary Citation Related Structures:
1QNX - PubMed Abstract:
Ves v 5 is one of three major allergens found in yellow-jacket venom: phospholipase A(1) (Ves v 1), hyaluronidase (Ves v 2), and antigen 5 (Ves v 5). Ves v 5 is related by high amino acid sequence identity to pathogenesis-related proteins including proteins from mammals, reptiles, insects, fungi, and plants. The crystal structure of Ves v 5 has been solved and refined to a resolution of 1.9 A. The majority of residues conserved between the pathogenesis-related proteins can be rationalized in terms of hydrogen bonding patterns and hydrophobic interactions defining an alpha-beta-alpha sandwich core structure. A small number of consensus residues are solvent exposed (including two adjacent histidines) and located in an elongated cavity that forms a putative active site. The site has no structural resemblance to previously characterized enzymes. Homologous antigen 5's from a large number of different yellow jackets, hornets, and paper wasps are known and patients show varying extents of cross-reactivity to the related antigen 5's. The structure of Ves v 5 allows a detailed analysis of the epitopes that may participate in antigenic cross-reactivity, findings that are useful for the development of a vaccine for treatment of insect allergy.
- Protein Structure Group, Department of Chemistry, University of Copenhagen, Copenhagen, Denmark. anette@crc.dk
Organizational Affiliation: