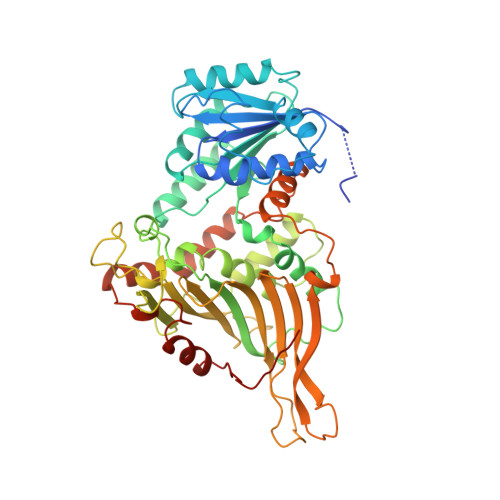
Human Glucose-6-Phosphate Dehydrogenase: The Crystal Structure Reveals a Structural Nadp+ Molecule and Provides Insights Into Enzyme Deficiency
Au, S.W.N., Gover, S., Lam, V.M.S., Adams, M.J.(2000) Structure 8: 293
- PubMed: 10745013 Search on PubMed
- DOI: https://doi.org/10.1016/s0969-2126(00)00104-0
- Primary Citation Related Structures:
1QKI - PubMed Abstract:
Glucose-6-phosphate dehydrogenase (G6PD) catalyses the first committed step in the pentose phosphate pathway; the generation of NADPH by this enzyme is essential for protection against oxidative stress. The human enzyme is in a dimer<-->tetramer equilibrium and its stability is dependent on NADP(+) concentration. G6PD deficiency results from many different point mutations in the X-linked gene encoding G6PD and is the most common human enzymopathy. Severe deficiency causes chronic non-spherocytic haemolytic anaemia; the usual symptoms are neonatal jaundice, favism and haemolytic anaemia. We have determined the first crystal structure of a human G6PD (the mutant Canton, Arg459-->Leu) at 3 A resolution. The tetramer is a dimer of dimers. Despite very similar dimer topology, there are two major differences from G6PD of Leuconostoc mesenteroides: a structural NADP(+) molecule, close to the dimer interface but integral to the subunit, is visible in all subunits of the human enzyme; and an intrasubunit disulphide bond tethers the otherwise disordered N-terminal segment. The few dimer-dimer contacts making the tetramer are charge-charge interactions. The importance of NADP(+) for stability is explained by the structural NADP(+) site, which is not conserved in prokaryotes. The structure shows that point mutations causing severe deficiency predominate close to the structural NADP(+) and the dimer interface, primarily affecting the stability of the molecule. They also indicate that a stable dimer is essential to retain activity in vivo. As there is an absolute requirement for some G6PD activity, residues essential for coenzyme or substrate binding are rarely modified.
- Laboratory of Molecular Biophysics, Department of Biochemistry, University of Oxford, The University of Hong Kong, Department of Biochemistry, Oxford, OX1 3QU, UK, Hong Kong.
Organizational Affiliation: