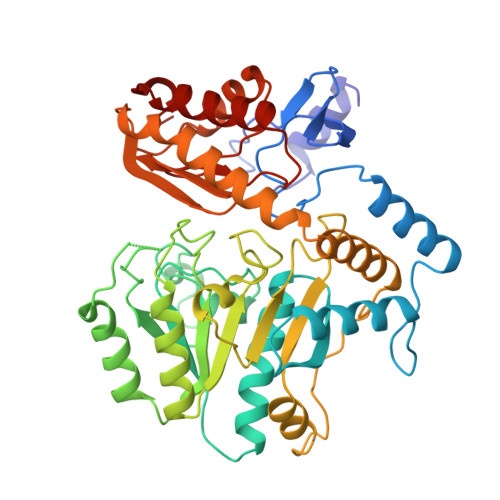
Crystal Structure of Diaminopelargonic Acid Synthase; Evolutionary Relationships between Pyridoxal-5'-Phosphate Dependent Enzymes
Kack, H., Sandmark, J., Gibson, K.J., Schneider, G., Lindqvist, Y.(1999) J Mol Biology 291: 857
- PubMed: 10452893 Search on PubMed
- DOI: https://doi.org/10.1006/jmbi.1999.2997
- Primary Citation Related Structures:
1QJ3, 1QJ5 - PubMed Abstract:
The three-dimensional structure of diaminopelargonic acid synthase, a vitamin B6-dependent enzyme in the pathway of the biosynthesis of biotin, has been determined to 1.8 A resolution by X-ray crystallography. The structure was solved by multi-wavelength anomalous diffraction techniques using a crystal derivatized with mercury ions. The protein model has been refined to a crystallographic R -value of 17.5% (R -free 22.6%). Each enzyme subunit consists of two domains, a large domain (residues 50-329) containing a seven-stranded predominantly parallel beta-sheet, surrounded by alpha-helices, and a small domain comprising residues 1-49 and 330-429. Two subunits, related by a non-crystallographic dyad in the crystals, form the homodimeric molecule, which contains two equal active sites. Pyridoxal-5'-phosphate is bound in a cleft formed by both domains of one subunit and the large domain of the second subunit. The cofactor is anchored to the enzyme by a covalent linkage to the side-chain of the invariant residue Lys274. The phosphate group interacts with main-chain nitrogen atoms and the side-chain of Ser113, located at the N terminus of an alpha-helix. The pyridine nitrogen forms a hydrogen bond to the side-chain of the invariant residue Asp245. Electron density corresponding to a metal ion, most likely Na(+), was found in a tight turn at the surface of the enzyme. Structure analysis reveals that diaminopelargonic acid synthase belongs to the family of vitamin B6-dependent aminotransferases with the same fold as originally observed in aspartate aminotransferase. A multiple structure alignment of enzymes in this family indicated that they form at least six different subclasses. Striking differences in the fold of the N-terminal part of the polypeptide chain are one of the hallmarks of these subclasses. Diaminopelargonic acid synthase is a member of the aminotransferase subclass III. From the structure of the non-productive complex of the holoenzyme with the substrate 7-keto-8-aminopelargonic acid the location of the active site and residues involved in substrate binding have been identified.
- Division of Structural Biology Department of Medical Biochemistry and Biophysics, Karolinska Institutet, Doktorsringen 9, Stockholm, S-171 77, Sweden.
Organizational Affiliation: