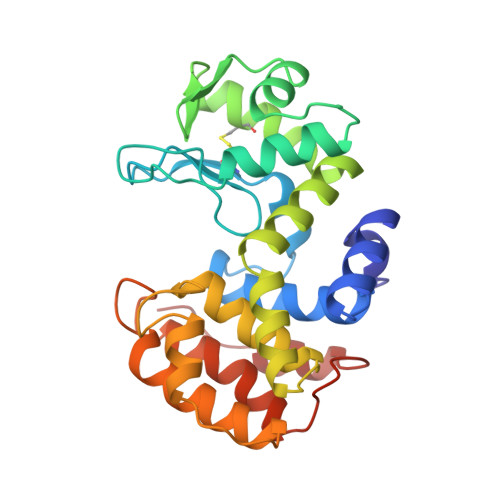
Crystal structure of chitosanase from Bacillus circulans MH-K1 at 1.6-A resolution and its substrate recognition mechanism.
Saito, J., Kita, A., Higuchi, Y., Nagata, Y., Ando, A., Miki, K.(1999) J Biological Chem 274: 30818-30825
- PubMed: 10521473 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.274.43.30818
- Primary Citation Related Structures:
1QGI - PubMed Abstract:
Chitosanase from Bacillus circulans MH-K1 is a 29-kDa extracellular protein composed of 259 amino acids. The crystal structure of chitosanase from B. circulans MH-K1 has been determined by multiwavelength anomalous diffraction method and refined to crystallographic R = 19.2% (R(free) = 23.5%) for the diffraction data at 1.6-A resolution collected by synchrotron radiation. The enzyme has two globular upper and lower domains, which generate the active site cleft for the substrate binding. The overall molecular folding is similar to chitosanase from Streptomyces sp. N174, although there is only 20% identity at the amino acid sequence level between both chitosanases. However, there are three regions in which the topology is remarkably different. In addition, the disulfide bridge between Cys(50) and Cys(124) joins the beta1 strand and the alpha7 helix, which is not conserved among other chitosanases. The orientation of two backbone helices, which connect the two domains, is also different and is responsible for the differences in size and shape of the active site cleft in these two chitosanases. This structural difference in the active site cleft is the reason why the enzymes specifically recognize different substrates and catalyze different types of chitosan degradation.
- Department of Chemistry, Graduate School of Science, Kyoto University, Sakyo-ku, Kyoto 606-8502, Japan.
Organizational Affiliation: