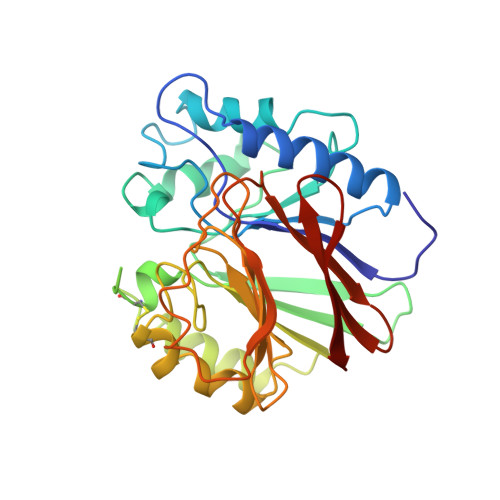
Crystal structure of a mammalian purple acid phosphatase.
Uppenberg, J., Lindqvist, F., Svensson, C., Ek-Rylander, B., Andersson, G.(1999) J Mol Biology 290: 201-211
- PubMed: 10388567 Search on PubMed
- DOI: https://doi.org/10.1006/jmbi.1999.2896
- Primary Citation Related Structures:
1QFC - PubMed Abstract:
Tartrate-resistant acid phosphatase (TRAP) is a mammalian di-iron- containing enzyme that belongs to the family of purple acid phosphatases (PAP). It is highly expressed in a limited number of tissues, predominantly in bone-resorbing osteoclasts and in macrophages of spleen. We have determined the crystal structure of rat TRAP in complex with a phosphate ion to 2.7 A resolution. The fold resembles that of the catalytic domain of kidney bean purple acid phosphatase (KBPAP), although the sequence similarity is limited to the active site residues. A surface loop near the active site is absent due to proteolysis, leaving the active-site easily accessible from the surrounding solvent. This, we believe, gives a structural explanation for the observed proteolytic activation of TRAP. The current structure was determined at a relatively high pH and without any external reducing agents. It is likely that it represents an oxidized and therefore catalytically inactive form of the enzyme.
- Department of Structural Chemistry, Pharmacia and Upjohn, Lindhagensgatan 133, Stockholm, S-112 87, Sweden. jonas.uppenberg@eu.pnu.com
Organizational Affiliation: