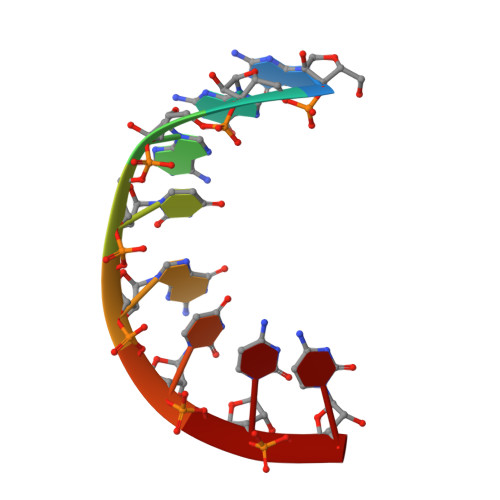
Investigation of the structural basis for thermodynamic stabilities of tandem GU wobble pairs: NMR structures of (rGGAGUUCC)2 and (rGGAUGUCC)2.
McDowell, J.A., He, L., Chen, X., Turner, D.H.(1997) Biochemistry 36: 8030-8038
- PubMed: 9201950 Search on PubMed
- DOI: https://doi.org/10.1021/bi970122c
- Primary Citation Related Structures:
1QES, 1QET - PubMed Abstract:
The symmetric, tandem GU mismatch motifs, and , which only differ in the mismatch order, have an average difference in thermodynamic stability of 2 kcal/mol at 37 degrees C. Thermodynamic studies of duplexes containing these motifs indicate the effect is largely localized to the mismatches and adjacent base pairs. The three-dimensional structures of two representative duplexes, (rGGAGUUCC)2 and (rGGAUGUCC)2, were determined by two-dimensional NMR and a simulated annealing protocol. Local deviations are similar to other intrahelical GU mismatches with little effect on backbone torsion angles and a slight overtwisting between the base pair 5' of the G of the mismatch and the mismatch itself. Comparisons of the resulting stacking patterns along with electrostatic potential maps suggest that interactions between highly negative electrostatic regions between base pairs may play a role in the observed thermodynamic differences.
- Department of Chemistry, University of Rochester, Rochester, New York 14627-0216, USA.
Organizational Affiliation: