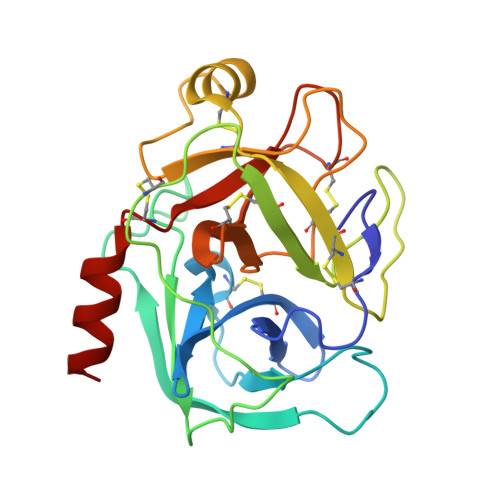
Crystallographic analysis of potent and selective factor Xa inhibitors complexed to bovine trypsin.
Whitlow, M., Arnaiz, D.O., Buckman, B.O., Davey, D.D., Griedel, B., Guilford, W.J., Koovakkat, S.K., Liang, A., Mohan, R., Phillips, G.B., Seto, M., Shaw, K.J., Xu, W., Zhao, Z., Light, D.R., Morrissey, M.M.(1999) Acta Crystallogr D Biol Crystallogr 55: 1395-1404
- PubMed: 10417407 Search on PubMed
- DOI: https://doi.org/10.1107/s0907444999007350
- Primary Citation Related Structures:
1QA0, 1QB1, 1QB6, 1QB9, 1QBN, 1QBO - PubMed Abstract:
Factor Xa is a serine protease which activates thrombin (factor IIa) and plays a key regulatory role in the blood-coagulation cascade. Factor Xa is, therefore, an important target for the design of anti-thrombotics. Both factor Xa and thrombin share sequence and structural homology with trypsin. As part of a factor Xa inhibitor-design program, a number of factor Xa inhibitors were crystallographically studied complexed to bovine trypsin. The structures of one diaryl benzimidazole, one diaryl carbazole and three diaryloxypyridines are described. All five compounds bind to trypsin in an extended conformation, with an amidinoaryl group in the S1 pocket and a second basic/hydrophobic moiety bound in the S4 pocket. These binding modes all bear a resemblance to the reported binding mode of DX-9065a in bovine trypsin and human factor Xa.
- Berlex Biosciences, 15049 San Pablo Avenue, PO Box 4099, Richmond, California 94804, USA. marc_whitlow@berlex.com
Organizational Affiliation: