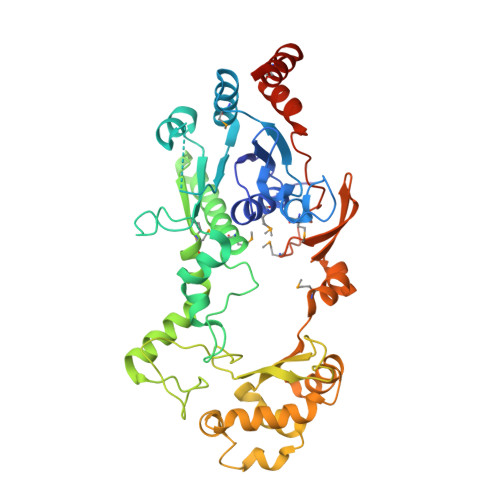
Structure of Escherichia coli YfdW, a type III CoA transferase.
Gogos, A., Gorman, J., Shapiro, L.(2004) Acta Crystallogr D Biol Crystallogr 60: 507-511
- PubMed: 14993676 Search on PubMed
- DOI: https://doi.org/10.1107/S0907444904000034
- Primary Citation Related Structures:
1PQY, 1Q6Y, 1Q7E - PubMed Abstract:
Crystal structures are reported for free and coenzyme A (CoA) bound forms of the YfdW protein from Escherichia coli, a representative type III CoA transferase. The structures reveal a two-domain protomer with interdomain connections forming a ring-like structure with a large central hole. Two protomers associate to form a highly intertwined dimer in which the hole of each ring is filled by the partner molecule. Each protomer binds a single CoA molecule and these CoA-binding sites are distant from one another in the dimer.
- Department of Biochemistry and Molecular Biophysics, Columbia University College of Physicians and Surgeons, 630 West 168th Street, New York, NY 10032, USA.
Organizational Affiliation: