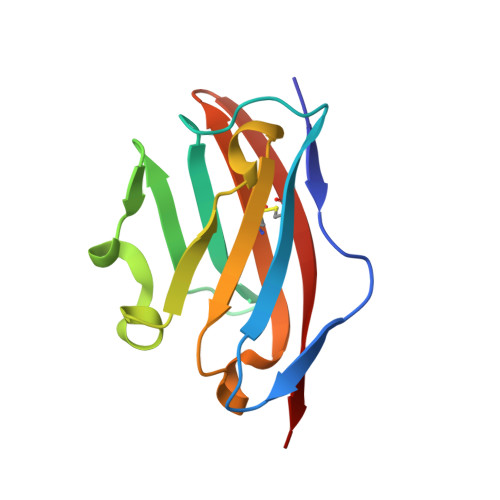
The crystal structure of myelin oligodendrocyte glycoprotein, a key autoantigen in multiple sclerosis
Clements, C.S., Reid, H.H., Beddoe, T., Tynan, F.E., Perugini, M.A., Johns, T.G., Bernard, C.C., Rossjohn, J.(2003) Proc Natl Acad Sci U S A 100: 11059-11064
- PubMed: 12960396 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1833158100
- Primary Citation Related Structures:
1PY9 - PubMed Abstract:
Myelin oligodendrocyte glycoprotein (MOG) is a key CNS-specific autoantigen for primary demyelination in multiple sclerosis. Although the disease-inducing role of MOG has been established, its precise function in the CNS remains obscure. To gain new insights into the physiological and immunopathological role of MOG, we determined the 1.8-A crystal structure of the MOG extracellular domain (MOGED). MOGED adopts a classical Ig (Ig variable domain) fold that was observed to form an antiparallel head-to-tail dimer. A dimeric form of native MOG was observed, and MOGED was also shown to dimerize in solution, consistent with the view of MOG acting as a homophilic adhesion receptor. The MOG35-55 peptide, a major encephalitogenic determinant recognized by both T cells and demyelinating autoantibodies, is partly occluded within the dimer interface. The structure of this key autoantigen suggests a relationship between the dimeric form of MOG within the myelin sheath and a breakdown of immunological tolerance to MOG that is observed in multiple sclerosis.
- Protein Crystallography Unit, Department of Biochemistry and Molecular Biology, School of Biomedical Sciences, Monash University, Clayton, Victoria 3168, Australia.
Organizational Affiliation: