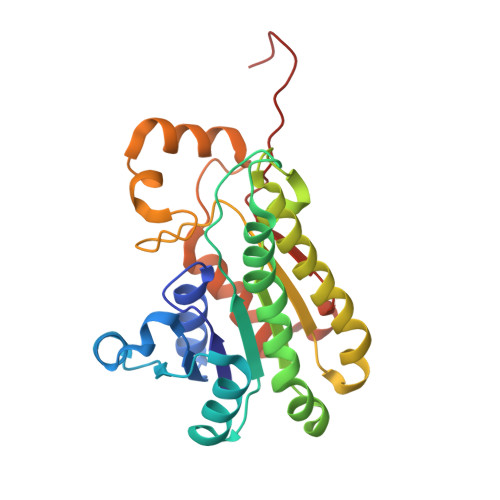
Structure and Mechanism of a Bacterial Haloalcohol Dehalogenase: a new variation of the short-chain dehydrogenase/reductase fold without an NAD(P)H binding site
de Jong, R.M., Tiesinga, J.J.W., Rozeboom, H.J., Kalk, K.H., Tang, L., Janssen, D.B., Dijkstra, B.W.(2003) EMBO J 22: 4933-4944
- PubMed: 14517233 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/emboj/cdg479
- Primary Citation Related Structures:
1PWX, 1PWZ, 1PX0 - PubMed Abstract:
Haloalcohol dehalogenases are bacterial enzymes that catalyze the cofactor-independent dehalogenation of vicinal haloalcohols such as the genotoxic environmental pollutant 1,3-dichloro-2-propanol, thereby producing an epoxide, a chloride ion and a proton. Here we present X-ray structures of the haloalcohol dehalogenase HheC from Agrobacterium radiobacter AD1, and complexes of the enzyme with an epoxide product and chloride ion, and with a bound haloalcohol substrate mimic. These structures support a catalytic mechanism in which Tyr145 of a Ser-Tyr-Arg catalytic triad deprotonates the haloalcohol hydroxyl function to generate an intramolecular nucleophile that substitutes the vicinal halogen. Haloalcohol dehalogenases are related to the widespread family of NAD(P)H-dependent short-chain dehydrogenases/reductases (SDR family), which use a similar Ser-Tyr-Lys/Arg catalytic triad to catalyze reductive or oxidative conversions of various secondary alcohols and ketones. Our results reveal the first structural details of an SDR-related enzyme that catalyzes a substitutive dehalogenation reaction rather than a redox reaction, in which a halide-binding site is found at the location of the NAD(P)H binding site. Structure-based sequence analysis reveals that the various haloalcohol dehalogenases have likely originated from at least two different NAD-binding SDR precursors.
- Department of Biophysical Chemistry, Groningen Biomolecular Sciences and Biotechnology Institute, University of Groningen, Nijenborgh 4, NL-9747 AG Groningen, The Netherlands.
Organizational Affiliation: