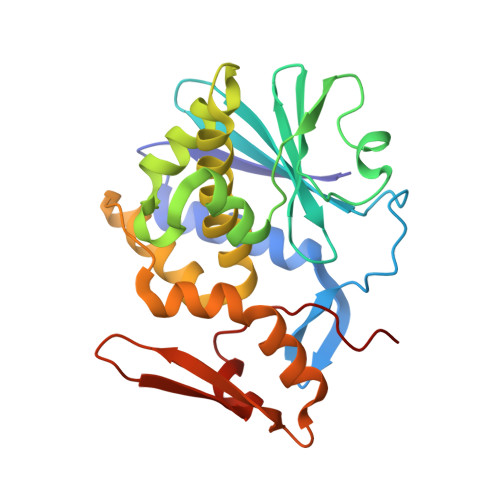
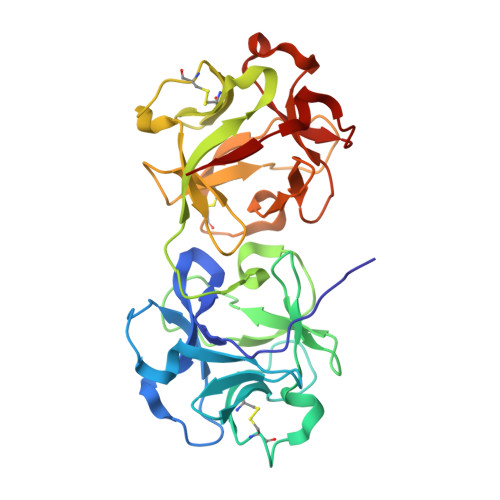
Mistletoe lectin I in complex with galactose and lactose reveals distinct sugar-binding properties
Mikeska, R., Wacker, R., Arni, R., Singh, T.P., Mikhailov, A., Gabdoulkhakov, A., Voelter, W., Betzel, C.(2005) Acta Crystallogr Sect F Struct Biol Cryst Commun 61: 17-25
- PubMed: 16508080 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S1744309104031501
- Primary Citation Related Structures:
1PUM, 1PUU - PubMed Abstract:
The structures of mistletoe lectin I (ML-I) from Viscum album complexed with lactose and galactose have been determined at 2.3 A resolution and refined to R factors of 20.9% (Rfree = 23.6%) and 20.9 (Rfree = 24.6%), respectively. ML-I is a heterodimer and belongs to the class of ribosome-inactivating proteins of type II, which consist of two chains. The A-chain has rRNA N-glycosidase activity and irreversibly inhibits eukaryotic ribosomes. The B-chain is a lectin and preferentially binds to galactose-terminated glycolipids and glycoproteins on cell membranes. Saccharide binding is performed by two binding sites in subdomains alpha1 and gamma2 of the ML-I B-chain separated by approximately 62 A from each other. The favoured binding of galactose in subdomain alpha1 is achieved via hydrogen bonds connecting the 4-hydroxyl and 3-hydroxyl groups of the sugar moiety with the side chains of Asp23B, Gln36B and Lys41B and the main chain of 26B. The aromatic ring of Trp38B on top of the preferred binding pocket supports van der Waals packing of the apolar face of galactose and stabilizes the sugar-lectin complex. In the galactose-binding site II of subdomain gamma2, Tyr249B provides the hydrophobic stacking and the side chains of Asp235B, Gln238B and Asn256B are hydrogen-bonding partners for galactose. In the case of the galactose-binding site I, the 2-hydroxyl group also stabilizes the sugar-protein complex, an interaction thus far rarely detected in galactose-specific lectins. Finally, a potential third low-affinity galactose-binding site in subunit beta1 was identified in the present ML-I structures, in which a glycerol molecule from the cryoprotectant buffer has bound, mimicking the sugar compound.
- Institute of Biochemistry and Food Chemistry, University of Hamburg, c/o DESY, Notkestrasse 85, 22603 Hamburg, Germany.
Organizational Affiliation: