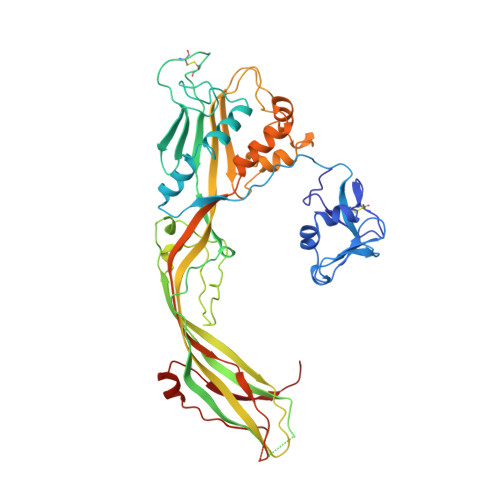
Structure of the Aeromonas toxin proaerolysin in its water-soluble and membrane-channel states.
Parker, M.W., Buckley, J.T., Postma, J.P., Tucker, A.D., Leonard, K., Pattus, F., Tsernoglou, D.(1994) Nature 367: 292-295
- PubMed: 7510043 Search on PubMed
- DOI: https://doi.org/10.1038/367292a0
- Primary Citation Related Structures:
1PRE - PubMed Abstract:
Aerolysin is chiefly responsible for the pathogenicity of Aeromonas hydrophila, a bacterium associated with diarrhoeal diseases and deep wound infections. Like many other microbial toxins, the protein changes in a multistep process from a completely water-soluble form to produce a transmembrane channel that destroys sensitive cells by breaking their permeability barriers. Here we describe the structure of proaerolysin determined by X-ray crystallography at 2.8 A resolution. The protoxin (M(r) 52,000) adopts a novel protein fold. Images of an aerolysin oligomer derived from electron microscopy have assisted in constructing a model of the membrane channel and have led to the proposal of a scheme to account for insertion of the protein into lipid bilayers to form ion channels.
- St Vincent's Institute of Medical Research, Fitzroy, Victoria, Australia.
Organizational Affiliation: