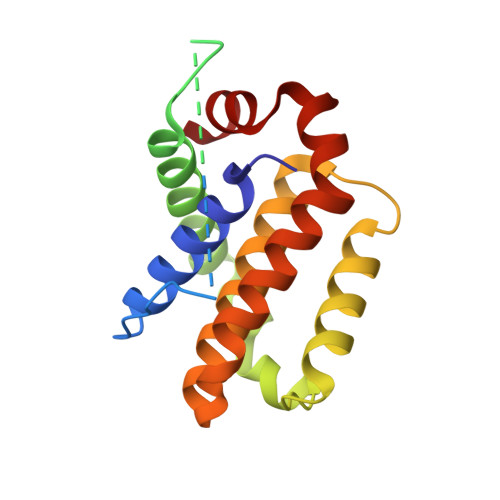
The structure of a Bcl-xl/Bim fragment complex: Implications for Bim function
Liu, X., Dai, S., Zhu, Y., Marrack, P., Kappler, J.W.(2003) Immunity 19: 341-352
- PubMed: 14499110 Search on PubMed
- DOI: https://doi.org/10.1016/s1074-7613(03)00234-6
- Primary Citation Related Structures:
1PQ0, 1PQ1 - PubMed Abstract:
After antigen-driven expansion, the majority of T cells involved in an immune response die rapidly by apoptosis dependent on the Bcl-2 related proteins, Bim and Bax or Bak. The details of how these proteins are activated and interact are still unclear. The crystal structure of mouse Bcl-x(L) bound to a long helical fragment of Bim indicates that the structure of Bim is very different from proteins with a Bcl-2-like fold and may leave the BH3 region of Bim constitutively exposed. Based on the structural homology between Bcl-x(L) and Bax, we predicted that binding of Bim to Bax would require displacement of the Bax penultimate alpha helix. Consistent with this prediction, truncation of this short helix was required for Bim/Bax interaction and led to spontaneous activation of Bax. Our results suggest a way in which both Bim and Bax/Bak might be required for activated T cell apoptosis.
- Howard Hughes Medical Institute, National Jewish Medical and Research Center, 1400 Jackson Street, Denver, CO 80206, USA.
Organizational Affiliation: