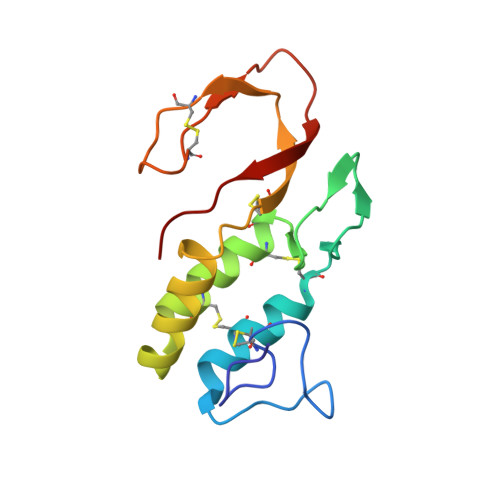
Crystal structure of bee-venom phospholipase A2 in a complex with a transition-state analogue.
Scott, D.L., Otwinowski, Z., Gelb, M.H., Sigler, P.B.(1990) Science 250: 1563-1566
- PubMed: 2274788 Search on PubMed
- DOI: https://doi.org/10.1126/science.2274788
- Primary Citation Related Structures:
1POC - PubMed Abstract:
The 2.0 angstroms crystal structure of a complex containing bee-venom phospholipase A2 (PLA2) and a phosphonate transition-state analogue was solved by multiple isomorphous replacement. The electron-density map is sufficiently detailed to visualize the proximal sugars of the enzyme's N-linked carbohydrate and a single molecule of the transition-state analogue bound ot its active center. Although bee-venom PLA2 does not belong to the large homologous Class I/II family that encompasses most other well-studied PLA2s, there is segmental sequence similarity and conservation of many functional substructures. Comparison of the bee-venom enzyme with other phospholipase structures provides compelling evidence for a common catalytic mechanism.
- Department of Molecular Biophysics and Biochemistry, Yale University, New Haven, CT 06511.
Organizational Affiliation: