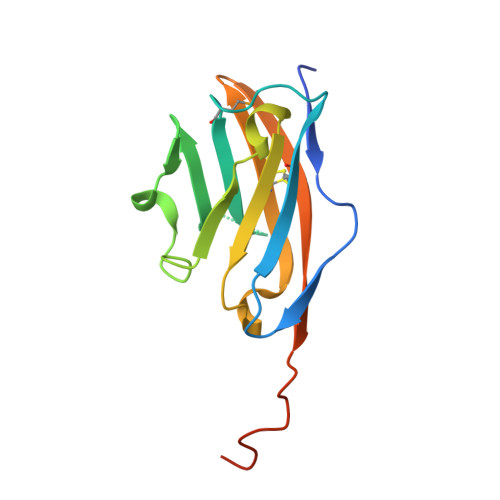
Structural insights into the antigenicity of myelin oligodendrocyte glycoprotein
Breithaupt, C., Schubart, A., Zander, H., Skerra, A., Huber, R., Linington, C., Jacob, U.(2003) Proc Natl Acad Sci U S A 100: 9446-9451
- PubMed: 12874380 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1133443100
- Primary Citation Related Structures:
1PKO, 1PKQ - PubMed Abstract:
Multiple sclerosis is a chronic disease of the central nervous system (CNS) characterized by inflammation, demyelination, and axonal loss. The immunopathogenesis of demyelination in multiple sclerosis involves an autoantibody response to myelin oligodendrocyte glycoprotein (MOG), a type I transmembrane protein located at the surface of CNS myelin. Here we present the crystal structures of the extracellular domain of MOG (MOGIgd) at 1.45-A resolution and the complex of MOGIgd with the antigen-binding fragment (Fab) of the MOG-specific demyelinating monoclonal antibody 8-18C5 at 3.0-A resolution. MOGIgd adopts an IgV like fold with the A'GFCC'C" sheet harboring a cavity similar to the one used by the costimulatory molecule B7-2 to bind its ligand CTLA4. The antibody 8-18C5 binds to three loops located at the membrane-distal side of MOG with a surprisingly dominant contribution made by MOG residues 101-108 containing a strained loop that forms the upper edge of the putative ligand binding site. The sequence R101DHSYQEE108 is unique for MOG, whereas large parts of the remaining sequence are conserved in potentially tolerogenic MOG homologues expressed outside the immuno-privileged environment of the CNS. Strikingly, the only sequence identical to DHSYQEE was found in a Chlamydia trachomatis protein of unknown function, raising the possibility that Chlamydia infections may play a role in the MOG-specific autoimmune response in man. Our data provide the structural basis for the development of diagnostic and therapeutic strategies targeting the pathogenic autoantibody response to MOG.
- Abteilung Strukturforschung, Max-Planck-Institut für Biochemie, Am Klopferspitz 18a, 82152 Martinsried, Germany. breitha@biochem.mpg.de
Organizational Affiliation: