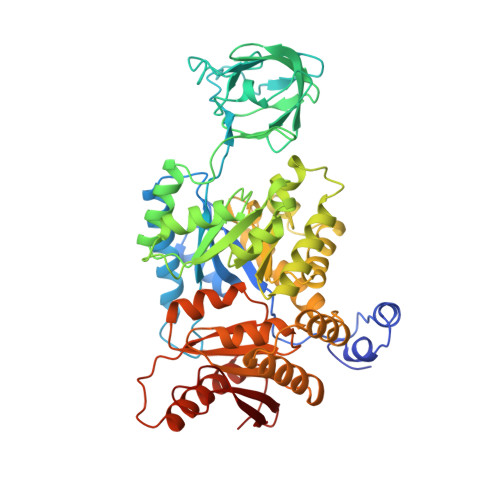
Refined three-dimensional structure of cat-muscle (M1) pyruvate kinase at a resolution of 2.6 A.
Allen, S.C., Muirhead, H.(1996) Acta Crystallogr D Biol Crystallogr 52: 499-504
- PubMed: 15299671 Search on PubMed
- DOI: https://doi.org/10.1107/S0907444995016040
- Primary Citation Related Structures:
1PKM - PubMed Abstract:
The three-dimensional structure of cat-muscle pyoruvate kinase has been refined at a resolution of 2.6 A. The details of the structure permit interpretation of the original heavy-atom studies and give insight into the importance of conserved residues in pyruvate kinases and the allosteric behaviour of the enzyme. There are a small number of essential residues which determine the relative orientations of domains and the precise nature of intersubunit contacts. Arginine residues are particularly important.
- Department of Biochemistry and Molecular Recognition Centre, School of Medical Sciences, University of Bristol.
Organizational Affiliation: