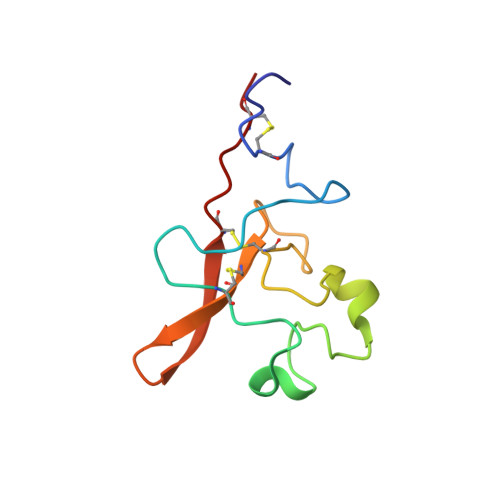
Solution structure of the tissue-type plasminogen activator kringle 2 domain complexed to 6-aminohexanoic acid an antifibrinolytic drug.
Byeon, I.J., Llinas, M.(1991) J Mol Biology 222: 1035-1051
- PubMed: 1762144 Search on PubMed
- DOI: https://doi.org/10.1016/0022-2836(91)90592-t
- Primary Citation Related Structures:
1PK2 - PubMed Abstract:
The solution structure of a recombinant tissue-type plasminogen activator kringle 2 domain, complexed with the antifibrinolytic drug 6-aminohexanoic acid (6-AHA) was determined via 1H nuclear magnetic resonance spectroscopy and dynamical simulated annealing calculations. The structure determination is based on 610 intramolecular kringle 2 and 14 intermolecular kringle 2-6-AHA interproton distance restraints, as well as on 82 torsion angle restraints. Three sets of simulated annealing structures were computed from three different classes of starting structures: (1) random conformations devoid of disulfide bridges; (2) random conformations that contain correct disulfide bonds; and (3) a folded conformation modeled after the homologous prothrombin kringle 1 X-ray crystallographic structure. All three sets of structures are well defined, with averaged atomic root-mean-square deviations between individual structures and mean set structures of 0.77, 0.99 and 0.70 A for backbone atoms, and 1.36, 1.55 and 1.41 A for all atoms, respectively. Kringle 2 is an oblate ellipsoid with overall dimensions of approximately 34 A x 30 A x 17 A. It exhibits a compact globular conformation characterized by a number of turns and loop elements as well as by one right-handed alpha-helix and five (1 extended and 4 rudimentary) antiparallel beta-sheets. The extended beta-sheet exhibits a right-handed twist. Close van der Waals' contacts between the Cys22-Cys63 and Cys51-Cys75 disulfide bridges and the central hydrophobic core composed of the Trp25, Leu46, His48a and Trp62 side-chains are among the distinguishing features of the kringle 2 fold. The binding site for 6-AHA appears as a rather exposed cleft with a negatively charged locus defined by the Asp55 and Asp57 side-chains, and with an aromatic pocket structured by the Tyr36, Trp62, His64 and Trp72 side-chains. The Trp62 and His64 rings line the back surface of the pocket, while the Tyr36 and Trp72 rings confine it from two sides. The Trp62 and Trp72 indole rings conform a V-shaped groove. The methyl groups of Val35 also contribute lipophilic character to the ligand-interacting surface. It is suggested that the positively charged side-chains of Lys34 and, potentially, Arg69 may favor interactions with the carboxylate group of the ligand. The Trp25 and Tyr74 aromatic rings, although conserved elements of the binding site structure, seem not to undergo direct contacts with the ligand.
- Department of Chemistry, Carnegie Mellon University, Pittsburgh, PA 15213.
Organizational Affiliation: