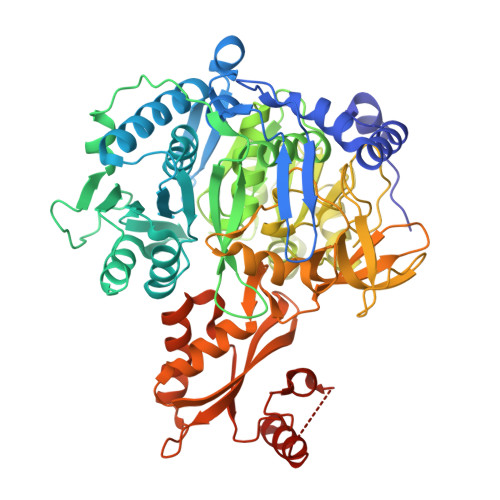
The 1.75 A Crystal Structure of Acetyl-CoA Synthetase Bound to Adenosine-5'-propylphosphate and Coenzyme A
Gulick, A.M., Starai, V.J., Horswill, A.R., Homick, K.M., Escalante-Semerena, J.C.(2003) Biochemistry 42: 2866-2873
- PubMed: 12627952 Search on PubMed
- DOI: https://doi.org/10.1021/bi0271603
- Primary Citation Related Structures:
1PG3, 1PG4 - PubMed Abstract:
Acetyl-coenzyme A synthetase catalyzes the two-step synthesis of acetyl-CoA from acetate, ATP, and CoA and belongs to a family of adenylate-forming enzymes that generate an acyl-AMP intermediate. This family includes other acyl- and aryl-CoA synthetases, firefly luciferase, and the adenylation domains of the modular nonribosomal peptide synthetases. We have determined the X-ray crystal structure of acetyl-CoA synthetase complexed with adenosine-5'-propylphosphate and CoA. The structure identifies the CoA binding pocket as well as a new conformation for members of this enzyme family in which the approximately 110-residue C-terminal domain exhibits a large rotation compared to structures of peptide synthetase adenylation domains. This domain movement presents a new set of residues to the active site and removes a conserved lysine residue that was previously shown to be important for catalysis of the adenylation half-reaction. Comparison of our structure with kinetic and structural data of closely related enzymes suggests that the members of the adenylate-forming family of enzymes may adopt two different orientations to catalyze the two half-reactions. Additionally, we provide a structural explanation for the recently shown control of enzyme activity by acetylation of an active site lysine.
- Hauptman-Woodward Medical Research Institute, Buffalo, New York 14203-1149, USA. gulick@hwi.buffalo.edu
Organizational Affiliation: