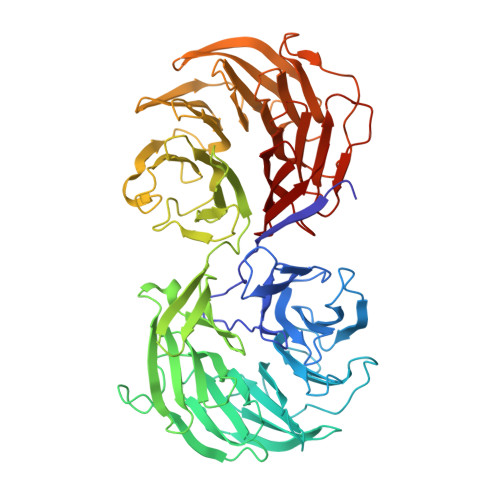
Identification of functional residues on Caenorhabditis elegans actin-interacting protein 1 (UNC-78) for disassembly of actin depolymerizing factor/cofilin-bound actin filaments.
Mohri, K., Vorobiev, S., Fedorov, A.A., Almo, S.C., Ono, S.(2004) J Biological Chem 279: 31697-31707
- PubMed: 15150269 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M403351200
- Primary Citation Related Structures:
1NR0, 1PEV - PubMed Abstract:
Actin-interacting protein 1 (AIP1) is a WD40 repeat protein that enhances actin filament disassembly in the presence of actin-depolymerizing factor (ADF)/cofilin. AIP1 also caps the barbed end of ADF/cofilin-bound actin filament. However, the mechanism by which AIP1 interacts with ADF/cofilin and actin is not clearly understood. We determined the crystal structure of Caenorhabditis elegans AIP1 (UNC-78), which revealed 14 WD40 modules arranged in two seven-bladed beta-propeller domains. The structure allowed for the mapping of conserved surface residues, and mutagenesis studies identified five residues that affected the ADF/cofilin-dependent actin filament disassembly activity. Mutations of these residues, which reside in blades 3 and 4 in the N-terminal propeller domain, had significant effects on the disassembly activity but did not alter the barbed end capping activity. These data support a model in which this conserved surface of AIP1 plays a direct role in enhancing fragmentation/depolymerization of ADF/cofilin-bound actin filaments but not in barbed end capping.
- Department of Pathology, Emory University, Atlanta, Georgia 30322, USA.
Organizational Affiliation: