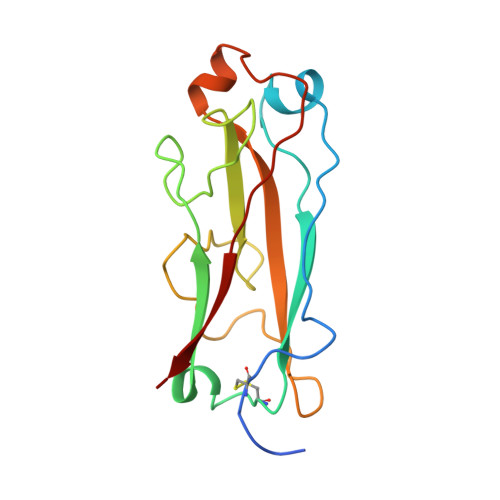
Structural basis of chaperone function and pilus biogenesis.
Sauer, F.G., Futterer, K., Pinkner, J.S., Dodson, K.W., Hultgren, S.J., Waksman, G.(1999) Science 285: 1058-1061
- PubMed: 10446050 Search on PubMed
- DOI: https://doi.org/10.1126/science.285.5430.1058
- Primary Citation Related Structures:
1PDK - PubMed Abstract:
Many Gram-negative pathogens assemble architecturally and functionally diverse adhesive pili on their surfaces by the chaperone-usher pathway. Immunoglobulin-like periplasmic chaperones escort pilus subunits to the usher, a large protein complex that facilitates the translocation and assembly of subunits across the outer membrane. The crystal structure of the PapD-PapK chaperone-subunit complex, determined at 2.4 angstrom resolution, reveals that the chaperone functions by donating its G(1) beta strand to complete the immunoglobulin-like fold of the subunit via a mechanism termed donor strand complementation. The structure of the PapD-PapK complex also suggests that during pilus biogenesis, every subunit completes the immunoglobulin-like fold of its neighboring subunit via a mechanism termed donor strand exchange.
- Department of Molecular Microbiology, Washington University School of Medicine, 660 South Euclid Avenue, St. Louis, MO 63110, USA.
Organizational Affiliation: