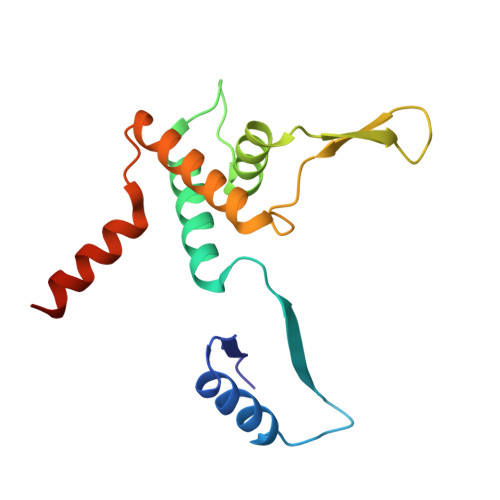
Functional similarities between phage lambda Orf and Escherichia coli RecFOR in initiation of genetic exchange
Maxwell, K.L., Reed, P., Zhang, R., Beasley, S., Walmsley, A.R., Curtis, F.A., Joachimiak, A., Edwards, A.M., Sharples, G.J.(2005) Proc Natl Acad Sci U S A 102: 11260-11265
- PubMed: 16076958 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.0503399102
- Primary Citation Related Structures:
1PC6 - PubMed Abstract:
Genetic recombination in bacteriophage lambda relies on DNA end processing by Exo to expose 3'-tailed strands for annealing and exchange by beta protein. Phage lambda encodes an additional recombinase, Orf, which participates in the early stages of recombination by supplying a function equivalent to the Escherichia coli RecFOR complex. These host enzymes assist loading of the RecA strand exchange protein onto ssDNA coated with ssDNA-binding protein. In this study, we purified the Orf protein, analyzed its biochemical properties, and determined its crystal structure at 2.5 angstroms. The homodimeric Orf protein is arranged as a toroid with a shallow U-shaped cleft, lined with basic residues, running perpendicular to the central cavity. Orf binds DNA, favoring single-stranded over duplex and with no obvious preference for gapped, 3'-tailed, or 5'-tailed substrates. An interaction between Orf and ssDNA-binding protein was indicated by far Western analysis. The functional similarities between Orf and RecFOR are discussed in relation to the early steps of recombinational exchange and the interplay between phage and bacterial recombinases.
- Centre for Infectious Diseases, Wolfson Research Institute, University of Durham, Queen's Campus, Stockton-on-Tees TS17 6BH, United Kingdom.
Organizational Affiliation: