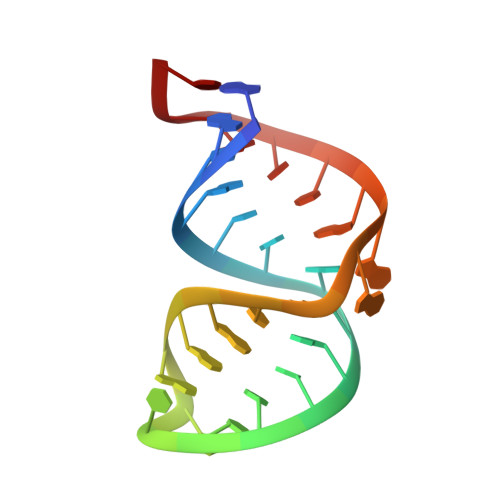
Structure of the A site of Escherichia coli 16S ribosomal RNA complexed with an aminoglycoside antibiotic.
Fourmy, D., Recht, M.I., Blanchard, S.C., Puglisi, J.D.(1996) Science 274: 1367-1371
- PubMed: 8910275 Search on PubMed
- DOI: https://doi.org/10.1126/science.274.5291.1367
- Primary Citation Related Structures:
1PBR - PubMed Abstract:
Aminoglycoside antibiotics that bind to 30S ribosomal A-site RNA cause misreading of the genetic code and inhibit translocation. The aminoglycoside antibiotic paromomycin binds specifically to an RNA oligonucleotide that contains the 30S subunit A site, and the solution structure of the RNA-paromomycin complex was determined by nuclear magnetic resonance spectroscopy. The antibiotic binds in the major groove of the model A-site RNA within a pocket created by an A-A base pair and a single bulged adenine. Specific interactions occur between aminoglycoside chemical groups important for antibiotic activity and conserved nucleotides in the RNA. The structure explains binding of diverse aminoglycosides to the ribosome, their specific activity against prokaryotic organisms, and various resistance mechanisms, and provides insight into ribosome function.
- Department of Chemistry and Biochemistry, Center for Molecular Biology of RNA, University of California, Santa Cruz, CA 95064, USA.
Organizational Affiliation: