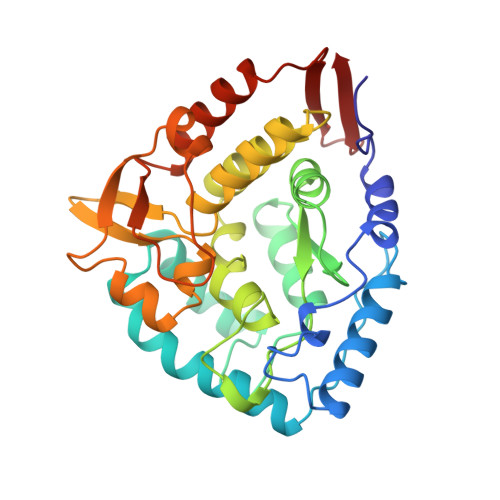
Crystal structure of the catalytic domain of human phenylalanine hydroxylase reveals the structural basis for phenylketonuria.
Erlandsen, H., Fusetti, F., Martinez, A., Hough, E., Flatmark, T., Stevens, R.C.(1997) Nat Struct Biol 4: 995-1000
- PubMed: 9406548 Search on PubMed
- DOI: https://doi.org/10.1038/nsb1297-995
- Primary Citation Related Structures:
1PAH - PubMed Abstract:
The 2.0 A crystal structure of the catalytic domain of human phenylalanine hydroxylase reveals a fold similar to that of tyrosine hydroxylase. It provides the first structural view of where mutations occur and a rationale to explain molecular mechanisms of the enzymatic phenotypes in the autosomal recessive disorder phenylketoneuria.