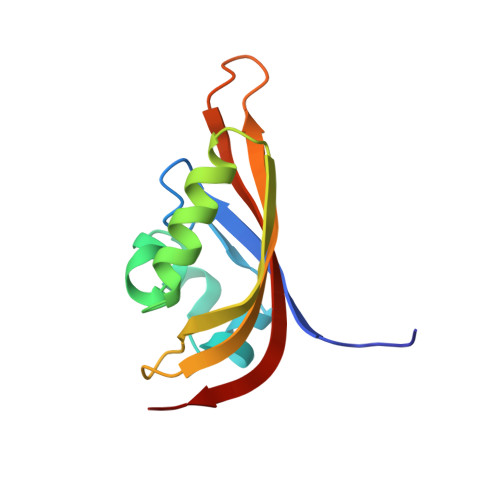
Structural basis for PAS domain heterodimerization in the basic helix-loop-helix-PAS transcription factor hypoxia-inducible factor.
Erbel, P.J., Card, P.B., Karakuzu, O., Bruick, R.K., Gardner, K.H.(2003) Proc Natl Acad Sci U S A 100: 15504-15509
- PubMed: 14668441 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.2533374100
- Primary Citation Related Structures:
1P97 - PubMed Abstract:
Biological responses to oxygen availability play important roles in development, physiological homeostasis, and many disease processes. In mammalian cells, this adaptation is mediated in part by a conserved pathway centered on the hypoxia-inducible factor (HIF). HIF is a heterodimeric protein complex composed of two members of the basic helix-loop-helix Per-ARNT-Sim (PAS) (ARNT, aryl hydrocarbon receptor nuclear translocator) domain family of transcriptional activators, HIFalpha and ARNT. Although this complex involves protein-protein interactions mediated by basic helix-loop-helix and PAS domains in both proteins, the role played by the PAS domains is poorly understood. To address this issue, we have studied the structure and interactions of the C-terminal PAS domain of human HIF-2alpha by NMR spectroscopy. We demonstrate that HIF-2alpha PAS-B binds the analogous ARNT domain in vitro, showing that residues involved in this interaction are located on the solvent-exposed side of the HIF-2alpha central beta-sheet. Mutating residues at this surface not only disrupts the interaction between isolated PAS domains in vitro but also interferes with the ability of full-length HIF to respond to hypoxia in living cells. Extending our findings to other PAS domains, we find that this beta-sheet interface is widely used for both intra- and intermolecular interactions, suggesting a basis of specificity and regulation of many types of PAS-containing signaling proteins.
- Departments of Biochemistry and Pharmacology, University of Texas Southwestern Medical Center, 5323 Harry Hines Boulevard, Dallas, TX 75390, USA.
Organizational Affiliation: