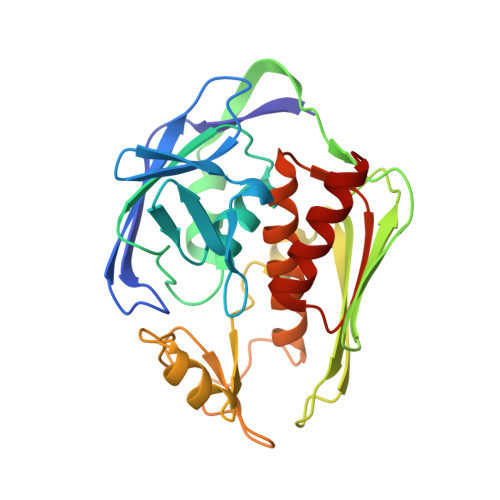
Crystal Structure of LpxC, a Zinc-Dependent Deacetylase Essential for Endotoxin Biosynthesis
Whittington, D.A., Rusche, K.M., Shin, H., Fierke, C.A., Christianson, D.W.(2003) Proc Natl Acad Sci U S A 100: 8146-8150
- PubMed: 12819349 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1432990100
- Primary Citation Related Structures:
1P42 - PubMed Abstract:
The outer leaflet of the outer membrane of the Gram-negative bacterium serves as a permeability barrier and is composed of lipopolysaccharide, also known as endotoxin. The membrane anchor of lipopolysaccharide is lipid A, the biosynthesis of which is essential for cell viability. The first committed step in lipid A biosynthesis is catalyzed by UDP-(3-O-(R-3-hydroxymyristoyl))-N-acetylglucosamine deacetylase (LpxC), a zinc-dependent deacetylase. Here we report the crystal structure of LpxC from Aquifex aeolicus, which reveals a new alpha+beta fold reflecting primordial gene duplication and fusion, as well as a new zinc-binding motif. The catalytic zinc ion resides at the base of an active-site cleft and adjacent to a hydrophobic tunnel occupied by a fatty acid. This tunnel accounts for the specificity of LpxC toward substrates and inhibitors bearing appropriately positioned 3-O-fatty acid substituents. Notably, simple inhibitors designed to target interactions in the hydrophobic tunnel bind with micromolar affinity, thereby representing a step toward the structure-based design of a potent, broad-spectrum antibacterial drug.
- Roy and Diana Vagelos Laboratories, Department of Chemistry, University of Pennsylvania, Philadelphia, PA 19104-6323, USA.
Organizational Affiliation: