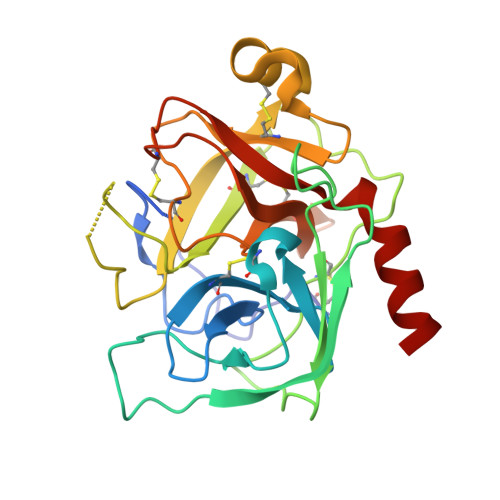
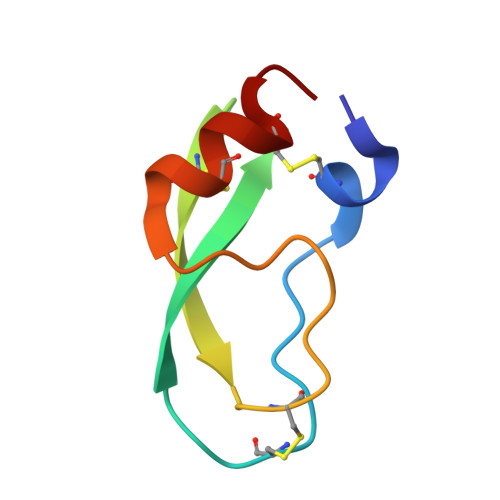
Structural consequences of accommodation of four non-cognate amino acid residues in the S1 pocket of bovine trypsin and chymotrypsin.
Helland, R., Czapinska, H., Leiros, I., Olufsen, M., Otlewski, J., Smalaas, A.O.(2003) J Mol Biology 333: 845-861
- PubMed: 14568540 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2003.08.059
- Primary Citation Related Structures:
1P2I, 1P2J, 1P2K, 1P2M, 1P2N, 1P2O, 1P2Q - PubMed Abstract:
Crystal structures of P1 Gly, Val, Leu and Phe bovine pancreatic trypsin inhibitor (BPTI) variants in complex with two serine proteinases, bovine trypsin and chymotrypsin, have been determined. The association constants for the four mutants with the two enzymes show that the enlargement of the volume of the P1 residue is accompanied by an increase of the binding energy, which is more pronounced for bovine chymotrypsin. Since the conformation of the P1 side-chains in the two S1 pockets is very similar, we suggest that the difference in DeltaG values between the enzymes must arise from the more polar environment of the S1 site of trypsin. This results mainly from the substitutions of Met192 and Ser189 observed in chymotrypsin with Gln192 and Asp189 present in trypsin. The more polar interior of the S1 site of trypsin is reflected by a much higher order of the solvent network in the empty pocket of the enzyme, as is observed in the complexes of the two enzymes with the P1 Gly BPTI variant. The more optimal binding of the large hydrophobic P1 residues by chymotrypsin is also reflected by shrinkage of the S1 pocket upon the accommodation of the cognate residues of this enzyme. Conversely, the S1 pocket of trypsin expands upon binding of such side-chains, possibly to avoid interaction with the polar residues of the walls. Further differentiation between the two enzymes is achieved by small differences in the shape of the S1 sites, resulting in an unequal steric hindrance of some of the side-chains, as observed for the gamma-branched P1 Leu variant of BPTI, which is much more favored by bovine chymotrypsin than trypsin. Analysis of the discrimination of beta-branched residues by trypsin and chymotrypsin is based on the complexes with the P1 Val BPTI variant. Steric repulsion of the P1 Val residue by the walls of the S1 pocket of both enzymes prevents the P1 Val side-chain from adopting the most optimal chi1 value.
- Norwegian Structural Biology Centre, Faculty of Science, University of Tromsø, 9037 Tromsø, Norway.
Organizational Affiliation: