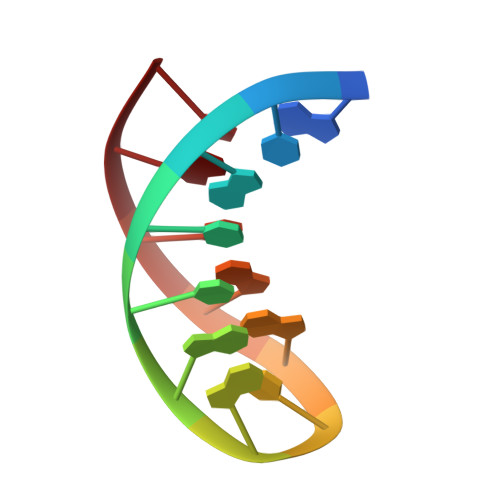
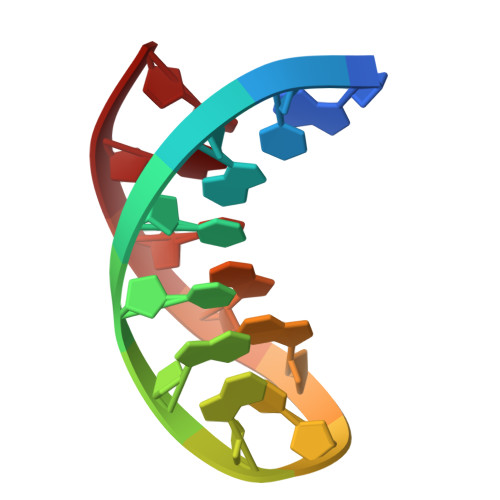
Sheared-type G(anti).C(syn) Base-pair: A Unique d(GXC) Loop Closure Motif.
Chin, K.H., Chou, S.H.(2003) J Mol Biol 329: 351-361
- PubMed: 12758081
- DOI: https://doi.org/10.1016/s0022-2836(03)00440-6
- Primary Citation of Related Structures:
1P0U - PubMed Abstract:
Stable DNA loop structures closed by a novel G.C base-pair have been determined for the single-residue d(GXC) loops (X=A, T, G or C) in low-salt solution by high-resolution nuclear magnetic resonance (NMR) techniques. The closing G.C base-pair in these loops is not of the canonical Watson-Crick type, but adopts instead a unique sheared-type (trans Watson-Crick/sugar-edge) pairing, like those occurring in the sheared mismatched G.A or A.C base-pair, to draw the two opposite strands together. The cytidine residue in the closing base-pair is transformed into the rare syn domain to form two H-bonds with the guanine base and to prevent the steric clash between the G 2NH(2) and the C H-5 protons. Besides, the sugar pucker of the syn cytidine is still located in the regular C2'-endo domain, unlike the C3'-endo domain adopted for the pyrimidines of the out-of-alternation left-handed Z-DNA structure. The facile formation of the compact d(GXC) loops closed by a unique sheared-type G(anti).C(syn) base-pair demonstrates the great potential of the single-stranded d(GXC) triplet repeats to fold into stable hairpins.
Organizational Affiliation:
Institute of Biochemistry, National Chung-Hsing University, Taichung, Taiwan, ROC.