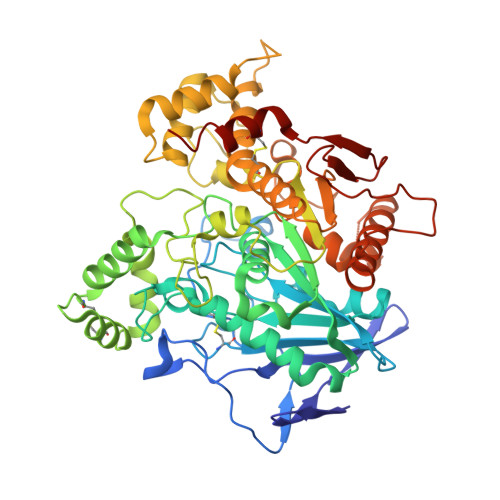
Crystal structure of human butyrylcholinesterase and of its complexes with substrate and products.
Nicolet, Y., Lockridge, O., Masson, P., Fontecilla-Camps, J.C., Nachon, F.(2003) J Biological Chem 278: 41141-41147
- PubMed: 12869558 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M210241200
- Primary Citation Related Structures:
1P0I, 1P0M, 1P0P, 1P0Q - PubMed Abstract:
Cholinesterases are among the most efficient enzymes known. They are divided into two groups: acetylcholinesterase, involved in the hydrolysis of the neurotransmitter acetylcholine, and butyrylcholinesterase of unknown function. Several crystal structures of the former have shown that the active site is located at the bottom of a deep and narrow gorge, raising the question of how substrate and products enter and leave. Human butyrylcholinesterase (BChE) has attracted attention because it can hydrolyze toxic esters such as cocaine or scavenge organophosphorus pesticides and nerve agents. Here we report the crystal structures of several recombinant truncated human BChE complexes and conjugates and provide a description for mechanistically relevant non-productive substrate and product binding. As expected, the structure of BChE is similar to a previously published theoretical model of this enzyme and to the structure of Torpedo acetylcholinesterase. The main difference between the experimentally determined BChE structure and its model is found at the acyl binding pocket that is significantly bigger than expected. An electron density peak close to the catalytic Ser(198) has been modeled as bound butyrate.
- Laboratoire de Cristallographie et Cristallogénèse des Protéines, Institut de Biologie Structurale "Jean-Pierre Ebel," CEA, UJF, CNRS, 41 rue Jules Horowitz, 38027 Grenoble Cedex 1, France.
Organizational Affiliation: