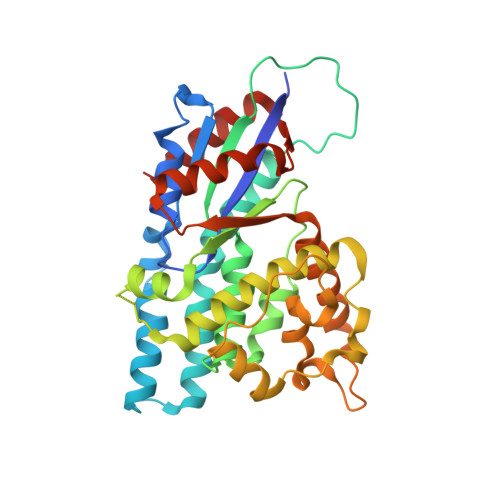
Crystal structure of varicella zoster virus thymidine kinase
Bird, L.E., Ren, J., Wright, A., Leslie, K.D., Degreve, B., Balzarini, J., Stammers, D.K.(2003) J Biological Chem 278: 24680-24687
- PubMed: 12686543 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M302025200
- Primary Citation Related Structures:
1OSN - PubMed Abstract:
Herpes virus thymidine kinases are responsible for the activation of nucleoside antiviral drugs including (E)-5-(2-bromovinyl)-2'-deoxyuridine. Such viral thymidine kinases (tk), beside having a broader substrate specificity compared with host cell enzymes, also show significant variation in nucleoside phosphorylation among themselves. We have determined the crystal structure of Varicella zoster virus (VZV, human herpes virus 3) thymidine kinase complexed with (E)-5-(2-bromovinyl)-2'-deoxyuridine 5'-monophosphate and ADP. Differences in the conformation of a loop region (residues 55-61) and the position of two alpha-helices at the subunit interface of VZV-tk compared with the herpes simplex virus type 1 (human herpes virus 1) enzyme give rise to changes in the positioning of residues such as tyrosine 66 and glutamine 90, which hydrogen bond to the substrate in the active site. Such changes in combination with the substitution in VZV-tk of two phenylalanine residues (in place of a tyrosine and methionine), which sandwich the substrate pyrimidine ring, cause an alteration in the positioning of the base. The interaction of the (E)-5-(2-bromovinyl)-2'-deoxyuridine deoxyribose ring with the protein is altered by substitution of tyrosine 21 and phenylalanine 139 (analagous to herpes simplex virus type 1 histidine 58 and tyrosine 172), which may explain some of the differences in nucleoside sugar selectivity between both enzymes. The altered active site architecture may also account for the differences in the substrate activity of ganciclovir for the two thymidine kinases. These data should be of use in the design of novel antiherpes and antitumor drugs.
- Division of Structural Biology, The Wellcome Trust Centre for Human Genetics, Henry Wellcome Building of Genomic Medicine, University of Oxford, Roosevelt Drive, Headington, United Kingdom.
Organizational Affiliation: