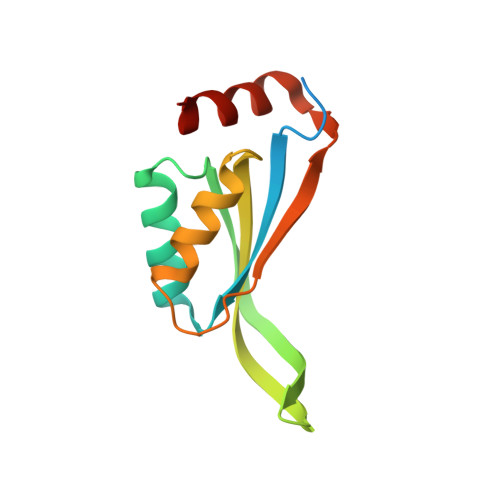
The Evolutionarily Conserved Trimeric Structure of CutA1 Proteins Suggests a Role in Signal Transduction
Arnesano, F., Banci, L., Benvenuti, M., Bertini, I., Calderone, V., Mangani, S., Viezzoli, M.S.(2003) J Biological Chem 278: 45999-46006
- PubMed: 12949080 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M304398200
- Primary Citation Related Structures:
1NAQ, 1OSC - PubMed Abstract:
CutA1 are a protein family present in bacteria, plants, and animals, including humans. Escherichia coli CutA1 is involved in copper tolerance, whereas mammalian proteins are implicated in the anchoring of acetylcholinesterase in neuronal cell membranes. The x-ray structures of CutA1 from E. coli and rat were determined. Both proteins are trimeric in the crystals and in solution through an inter-subunit beta-sheet formation. Each subunit consists of a ferredoxin-like (beta1alpha1beta2beta3alpha2beta4) fold with an additional strand (beta5), a C-terminal helix (alpha3), and an unusual extended beta-hairpin involving strands beta2 and beta3. The bacterial CutA1 is able to bind copper(II) in vitro through His2Cys coordination in a type II water-accessible site, whereas the rat protein precipitates in the presence of copper(II). The evolutionarily conserved trimeric assembly of CutA1 is reminiscent of the architecture of PII signal transduction proteins. This similarity suggests an intriguing role of CutA1 proteins in signal transduction through allosteric communications between subunits.
- Magnetic Resonance Center (CERM) and Department of Chemistry, University of Florence, Via Luigi Sacconi 6, 50019, Sesto Fiorentino, Florence, Italy.
Organizational Affiliation: