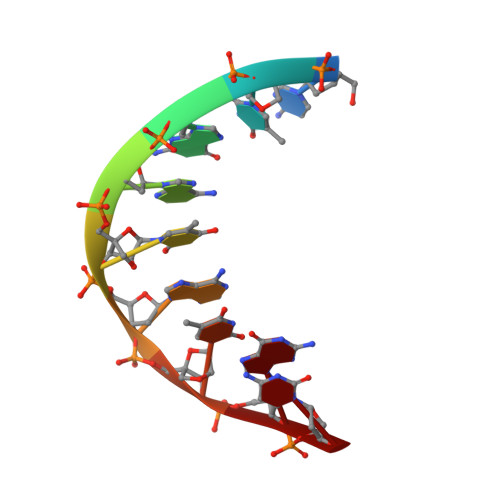
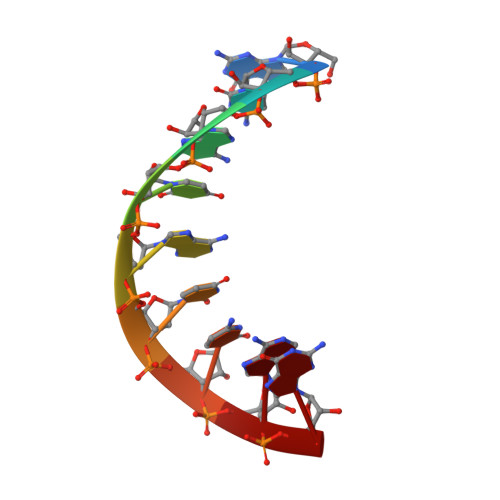
NMR Structure of an Alpha-L-Lna:RNA Hybrid: Structural Implications for Rnase H Recognition
Nielsen, J.T., Stein, P.C., Petersen, M.(2003) Nucleic Acids Res 31: 5858
- PubMed: 14530434 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gkg800
- Primary Citation Related Structures:
1OKF - PubMed Abstract:
Alpha-L-LNA (alpha-L-ribo configured locked nucleic acid) is a nucleotide analogue that raises the thermostability of nucleic acid duplexes by up to approximately 4 degrees C per inclusion. We have determined the NMR structure of a nonamer alpha-L-LNA:RNA hybrid with three alpha-L-LNA modifications. The geometry of this hybrid is intermediate between A- and B-type, all nucleobases partake in Watson-Crick base pairing and base stacking, and the global structure is very similar to that of the corresponding unmodified hybrid. The sugar-phosphate backbone is rearranged in the vicinity of the modified nucleotides. As a consequence, the phosphate groups following the modified nucleotides are rotated into the minor groove. It is interesting that the alpha-L-LNA:RNA hybrid, which has an elevation in melting temperature of 17 degrees C relative to the corresponding DNA:RNA hybrid, retains the global structure of this hybrid. To our knowledge, this is the first example of such a substantial increase in melting temperature of a nucleic acid analogue that does not act as an N-type (RNA) mimic. alpha-L-LNA:RNA hybrids are recognised by RNase H with subsequent cleavage of the RNA strand, albeit with slow rates. We attempt to rationalise this impaired enzyme activity from the rearrangement of the sugar-phosphate backbone of the alpha-L-LNA:RNA hybrid.
- Nucleic Acid Center, Department of Chemistry, University of Southern Denmark, 5230 Odense M, Denmark.
Organizational Affiliation: