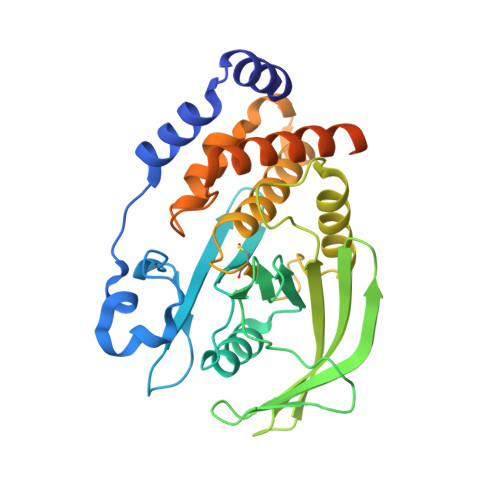
Oxidation state of the active-site cysteine in protein tyrosine phosphatase 1B.
van Montfort, R.L., Congreve, M., Tisi, D., Carr, R., Jhoti, H.(2003) Nature 423: 773-777
- PubMed: 12802339 Search on PubMed
- DOI: https://doi.org/10.1038/nature01681
- Primary Citation Related Structures:
1OES, 1OET, 1OEU, 1OEV - PubMed Abstract:
Protein tyrosine phosphatases regulate signal transduction pathways involving tyrosine phosphorylation and have been implicated in the development of cancer, diabetes, rheumatoid arthritis and hypertension. Increasing evidence suggests that the cellular redox state is involved in regulating tyrosine phosphatase activity through the reversible oxidization of the catalytic cysteine to sulphenic acid (Cys-SOH). But how further oxidation to the irreversible sulphinic (Cys-SO2H) and sulphonic (Cys-SO3H) forms is prevented remains unclear. Here we report the crystal structures of the regulatory sulphenic and irreversible sulphinic and sulphonic acids of protein tyrosine phosphatase 1B (PTP1B), an important enzyme in the negative regulation of the insulin receptor and a therapeutic target in type II diabetes and obesity. We also identify a sulphenyl-amide species that is formed through oxidation of its catalytic cysteine. Formation of the sulphenyl-amide causes large changes in the PTP1B active site, which are reversible by reduction with the cellular reducing agent glutathione. The sulphenyl-amide is a protective intermediate in the oxidative inhibition of PTP1B. In addition, it may facilitate reactivation of PTP1B by biological thiols and signal a unique state of the protein.
- Astex Technology Ltd, 436 Cambridge Science Park, Milton Road, Cambridge CB4 0QA, UK.
Organizational Affiliation: